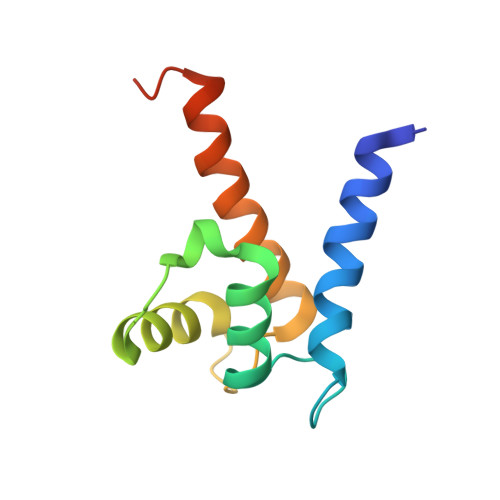
Phenothiazines inhibit S100A4 function by inducing protein oligomerization.
Malashkevich, V.N., Dulyaninova, N.G., Ramagopal, U.A., Liriano, M.A., Varney, K.M., Knight, D., Brenowitz, M., Weber, D.J., Almo, S.C., Bresnick, A.R.(2010) Proc Natl Acad Sci U S A 107: 8605-8610
- PubMed: 20421509 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0913660107
- Primary Citation Related Structures:
3KO0, 3M0W - PubMed Abstract:
S100A4, a member of the S100 family of Ca(2+)-binding proteins, regulates carcinoma cell motility via interactions with myosin-IIA. Numerous studies indicate that S100A4 is not simply a marker for metastatic disease, but rather has a direct role in metastatic progression. These observations suggest that S100A4 is an excellent target for therapeutic intervention. Using a unique biosensor-based assay, trifluoperazine (TFP) was identified as an inhibitor that disrupts the S100A4/myosin-IIA interaction. To examine the interaction of S100A4 with TFP, we determined the 2.3 A crystal structure of human Ca(2+)-S100A4 bound to TFP. Two TFP molecules bind within the hydrophobic target binding pocket of Ca(2+)-S100A4 with no significant conformational changes observed in the protein upon complex formation. NMR chemical shift perturbations are consistent with the crystal structure and demonstrate that TFP binds to the target binding cleft of S100A4 in solution. Remarkably, TFP binding results in the assembly of five Ca(2+)-S100A4/TFP dimers into a tightly packed pentameric ring. Within each pentamer most of the contacts between S100A4 dimers occurs through the TFP moieties. The Ca(2+)-S100A4/prochlorperazine (PCP) complex exhibits a similar pentameric assembly. Equilibrium sedimentation and cross-linking studies demonstrate the cooperative formation of a similarly sized S100A4/TFP oligomer in solution. Assays examining the ability of TFP to block S100A4-mediated disassembly of myosin-IIA filaments demonstrate that significant inhibition of S100A4 function occurs only at TFP concentrations that promote S100A4 oligomerization. Together these studies support a unique mode of inhibition in which phenothiazines disrupt the S100A4/myosin-IIA interaction by sequestering S100A4 via small molecule-induced oligomerization.
- Department of Biochemistry, Albert Einstein College of Medicine, Bronx, NY 10461, USA.
Organizational Affiliation: