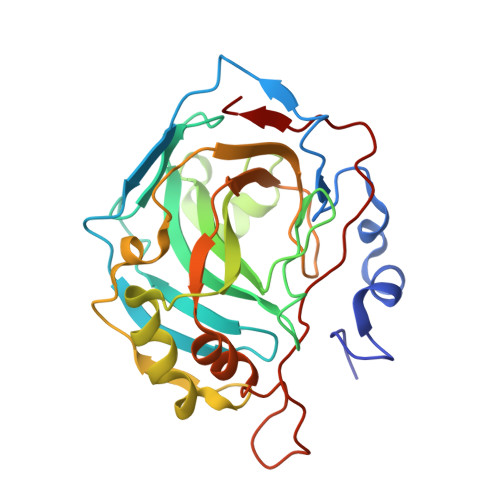
Neutron structure of human carbonic anhydrase II: implications for proton transfer.
Fisher, S.Z., Kovalevsky, A.Y., Domsic, J.F., Mustyakimov, M., McKenna, R., Silverman, D.N., Langan, P.A.(2010) Biochemistry 49: 415-421
- PubMed: 20025241 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi901995n
- Primary Citation Related Structures:
3KKX - PubMed Abstract:
Human carbonic anhydrase II (HCA II) catalyzes the reversible hydration of carbon dioxide to form bicarbonate and a proton. Despite many high-resolution X-ray crystal structures, mutagenesis, and kinetic data, the structural details of the active site, especially the proton transfer pathway, are unclear. A large HCA II crystal was prepared at pH 9.0 and subjected to vapor H-D exchange to replace labile hydrogens with deuteriums. Neutron diffraction studies were conducted at the Protein Crystallography Station at Los Alamos National Laboratory. The structure to 2.0 A resolution reveals several interesting active site features: (1) the Zn-bound solvent appearing to be predominantly a D(2)O molecule, (2) the orientation and hydrogen bonding pattern of solvent molecules in the active site cavity, (3) the side chain of His64 being unprotonated (neutral) and predominantly in an inward conformation pointing toward the zinc, and (4) the phenolic side chain of Tyr7 appearing to be unprotonated. The implications of these details are discussed, and a proposed mechanism for proton transfer is presented.
- Bioscience Division MS M888, Los Alamos National Laboratory, Los Alamos, New Mexico 87544, USA. zfisher@lanl.gov
Organizational Affiliation: