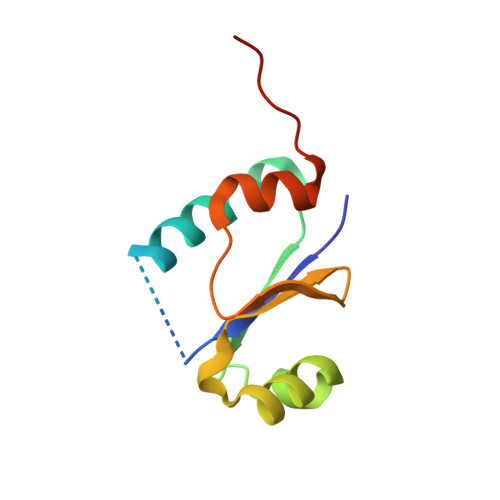
The 1.4 A crystal structure of the ArsD arsenic metallochaperone provides insights into its interaction with the ArsA ATPase.
Ye, J., Ajees, A.A., Yang, J., Rosen, B.P.(2010) Biochemistry 49: 5206-5212
- PubMed: 20507177 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi100571r
- Primary Citation Related Structures:
3KGK, 3MWH - PubMed Abstract:
Arsenic is a carcinogen that tops the Superfund list of hazardous chemicals. Bacterial resistance to arsenic is facilitated by ArsD, which delivers As(III) to the ArsA ATPase, the catalytic subunit of the ArsAB pump. Here we report the structure of the arsenic metallochaperone ArsD at 1.4 A and a model for its binding of metalloid. There are two ArsD molecules in the asymmetric unit. The overall structure of the ArsD monomer has a thioredoxin fold, with a core of four beta-strands flanked by four alpha-helices. Based on data from structural homologues, ArsD was modeled with and without bound As(III). ArsD binds one arsenic per monomer coordinated with the three sulfur atoms of Cys12, Cys13, and Cys18. Using this structural model, an algorithm was used to dock ArsD and ArsA. The resulting docking model provides testable predictions of the contact points of the two proteins and forms the basis for future experiments.
- Department of Biochemistry and Molecular Biology, Wayne State University School of Medicine, Detroit, Michigan 48201, USA.
Organizational Affiliation: