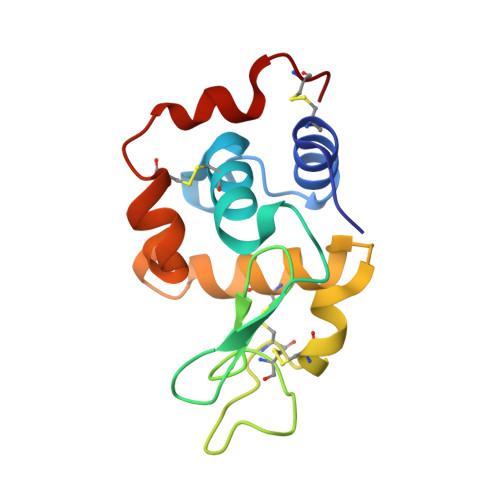
Specific derivatization of lysozyme in aqueous solution with Re(CO)3(H2O)3(+).
Binkley, S.L., Ziegler, C.J., Herrick, R.S., Rowlett, R.S.(2010) Chem Commun (Camb) 46: 1203-1205
- PubMed: 20449250 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1039/b923688k
- Primary Citation Related Structures:
3KAM - PubMed Abstract:
The reaction of Re(CO)(3)(H(2)O)(3)(+) with hen egg lysozyme in aqueous solution results in a single covalent adduct; single crystal X-ray diffraction shows that the rhenium tricarbonyl cation binds to His15 in two significantly populated rotamer conformations.
- Department of Chemistry, University of Akron, Akron, OH 44325-3601, USA.
Organizational Affiliation: