Mutational studies of the peptidoglycan hydrolase FlgJ of Sphingomonas sp. strain A1
Maruyama, Y., Ochiai, A., Itoh, T., Mikami, B., Hashimoto, W., Murata, K.(2010) J Basic Microbiol 50: 311-317
- PubMed: 20586063 Search on PubMed
- DOI: https://doi.org/10.1002/jobm.200900249
- Primary Citation Related Structures:
3K3T - PubMed Abstract:
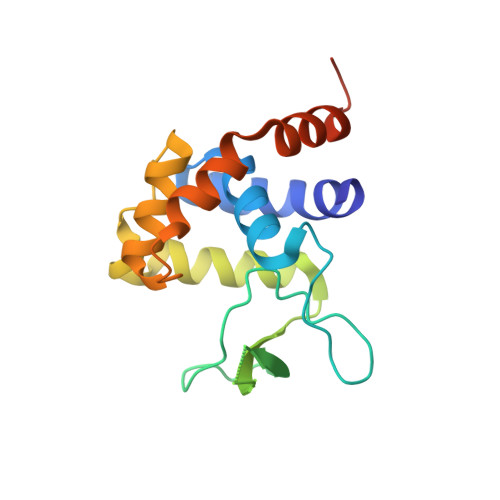
The flagellar protein FlgJ, a member of glycoside hydrolase family 73, has N- and C-terminal domains that are responsible for flagellar rod assembly and peptidoglycan hydrolysis, respectively. The crystal structure of the C-terminal domain of SPH1045 (SPH1045-C), the FlgJ from Sphingomonas sp. strain A1, showed a long cleft formed by two lobes, alpha and beta. In this study, seven site-specific mutants of residues in the cleft were prepared and analyzed. Enzyme activity was reduced most significantly in mutants E185A and Y281A, followed by E224A. A comparison of the crystal structure of the inactive mutant E185A with that of other related enzymes revealed that Glu185 is structurally reasonable as the proton donor and that Tyr281 is close to Glu185. Glu224 is, however, far from the catalytic site, which is inconsistent with the decreased activity exhibited by E224A. The structural flexibility of Glu224 and its neighboring residues observed in SPH1045-C may indicate that this region is able to change its conformation upon substrate binding.
- Laboratory of Basic and Applied Molecular Biotechnology, Graduate School of Agriculture, Kyoto University, Uji, Kyoto, Japan.
Organizational Affiliation: