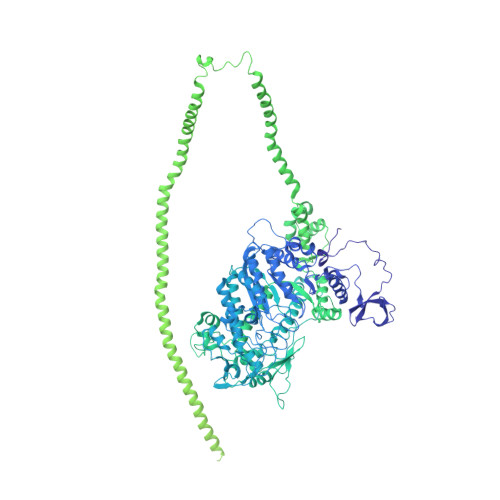
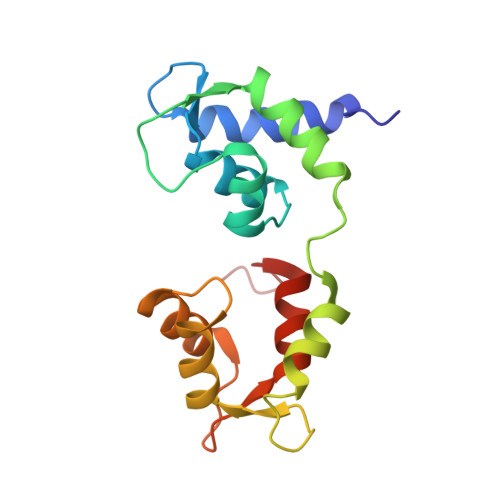
Conserved Intramolecular Interactions Maintain Myosin Interacting-Heads Motifs Explaining Tarantula Muscle Super-Relaxed State Structural Basis.
Alamo, L., Qi, D., Wriggers, W., Pinto, A., Zhu, J., Bilbao, A., Gillilan, R.E., Hu, S., Padron, R.(2016) J Mol Biology 428: 1142-1164
- PubMed: 26851071
- DOI: https://doi.org/10.1016/j.jmb.2016.01.027
- Primary Citation Related Structures:
3JBH - PubMed Abstract:
Tarantula striated muscle is an outstanding system for understanding the molecular organization of myosin filaments. Three-dimensional reconstruction based on cryo-electron microscopy images and single-particle image processing revealed that, in a relaxed state, myosin molecules undergo intramolecular head-head interactions, explaining why head activity switches off. The filament model obtained by rigidly docking a chicken smooth muscle myosin structure to the reconstruction was improved by flexibly fitting an atomic model built by mixing structures from different species to a tilt-corrected 2-nm three-dimensional map of frozen-hydrated tarantula thick filament. We used heavy and light chain sequences from tarantula myosin to build a single-species homology model of two heavy meromyosin interacting-heads motifs (IHMs). The flexibly fitted model includes previously missing loops and shows five intramolecular and five intermolecular interactions that keep the IHM in a compact off structure, forming four helical tracks of IHMs around the backbone. The residues involved in these interactions are oppositely charged, and their sequence conservation suggests that IHM is present across animal species. The new model, PDB 3JBH, explains the structural origin of the ATP turnover rates detected in relaxed tarantula muscle by ascribing the very slow rate to docked unphosphorylated heads, the slow rate to phosphorylated docked heads, and the fast rate to phosphorylated undocked heads. The conservation of intramolecular interactions across animal species and the presence of IHM in bilaterians suggest that a super-relaxed state should be maintained, as it plays a role in saving ATP in skeletal, cardiac, and smooth muscles.
- Centro de Biología Estructural, Instituto Venezolano de Investigaciones Científicas, Apartado 20632, Caracas 1020A, Venezuela. Electronic address: lorenzo.alamo@gmail.com.
Organizational Affiliation: