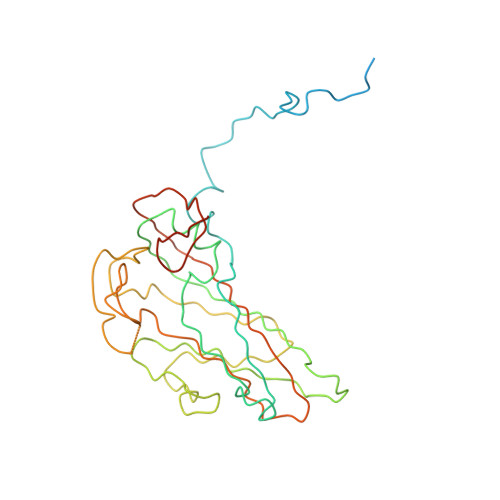
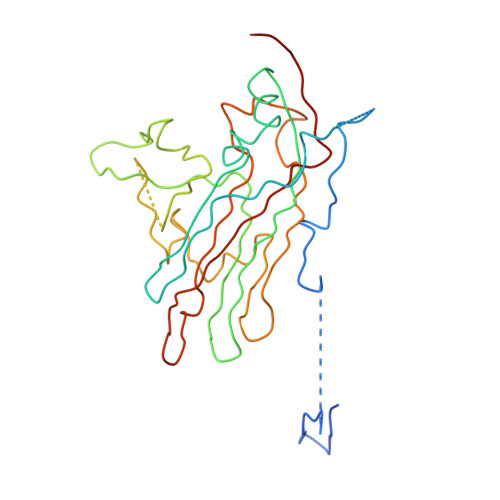
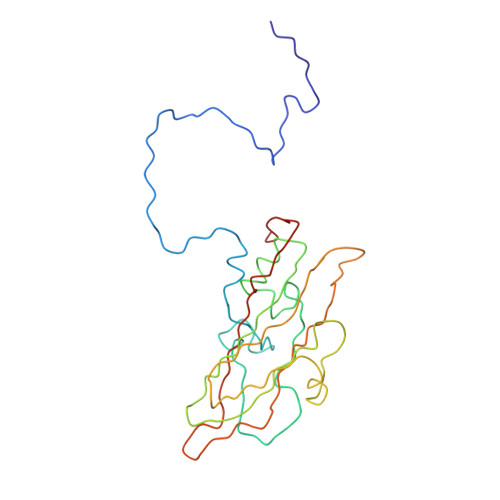
Cryo-Electron Microscopy Reconstruction Shows Poliovirus 135S Particles Poised for Membrane Interaction and RNA Release.
Butan, C., Filman, D.J., Hogle, J.M.(2014) J Virol 88: 1758-1770
- PubMed: 24257617 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.01949-13
- Primary Citation Related Structures:
3J48 - PubMed Abstract:
During infection, binding of mature poliovirus to cell surface receptors induces an irreversible expansion of the capsid, to form an infectious cell-entry intermediate particle that sediments at 135S. In these expanded virions, the major capsid proteins (VP1 to VP3) adopt an altered icosahedral arrangement to open holes in the capsid at 2-fold and quasi-3-fold axes, and internal polypeptides VP4 and the N terminus of VP1, which can bind membranes, become externalized. Cryo-electron microscopy images for 117,330 particles were collected using Leginon and reconstructed using FREALIGN. Improved rigid-body positioning of major capsid proteins established reliably which polypeptide segments become disordered or rearranged. The virus-to-135S transition includes expansion of 4%, rearrangements of the GH loops of VP3 and VP1, and disordering of C-terminal extensions of VP1 and VP2. The N terminus of VP1 rearranges to become externalized near its quasi-3-fold exit, binds to rearranged GH loops of VP3 and VP1, and attaches to the top surface of VP2. These details improve our understanding of subsequent stages of infection, including endocytosis and RNA transfer into the cytoplasm.
- Department of Biological Chemistry and Molecular Pharmacology, Harvard Medical School, Boston, Massachusetts, USA.
Organizational Affiliation: