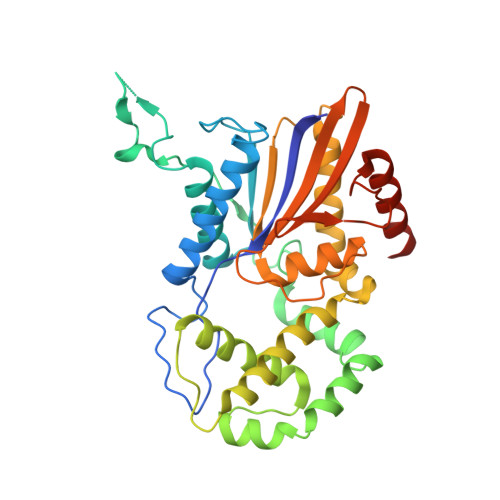
Crystal Structures of the histidine acid phosphatase from Francisella tularensis provide insight into substrate recognition.
Singh, H., Felts, R.L., Schuermann, J.P., Reilly, T.J., Tanner, J.J.(2009) J Mol Biology 394: 893-904
- PubMed: 19836403 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2009.10.009
- Primary Citation Related Structures:
3IT0, 3IT1, 3IT2, 3IT3 - PubMed Abstract:
Histidine acid phosphatases catalyze the transfer of a phosphoryl group from phosphomonoesters to water at acidic pH using an active-site histidine. The histidine acid phosphatase from the category A pathogen Francisella tularensis (FtHAP) has been implicated in intramacrophage survival and virulence, motivating interest in understanding the structure and mechanism of this enzyme. Here, we report a structure-based study of ligand recognition by FtHAP. The 1.70-A-resolution structure of FtHAP complexed with the competitive inhibitor l(+)-tartrate was solved using single-wavelength anomalous diffraction phasing. Structures of the ligand-free enzyme and the complex with inorganic phosphate were determined at resolutions of 1.85 and 1.70 A, respectively. The structure of the Asp261Ala mutant enzyme complexed with the substrate 3'-AMP was determined at 1.50 A resolution to gain insight into substrate recognition. FtHAP exhibits a two-domain fold similar to that of human prostatic acid phosphatase, consisting of an alpha/beta core domain and a smaller domain that caps the core domain. The structures show that the core domain supplies the phosphoryl binding site, catalytic histidine (His17), and an aspartic acid residue (Asp261) that protonates the leaving group, while the cap domain contributes residues that enforce substrate preference. FtHAP and human prostatic acid phosphatase differ in the orientation of the crucial first helix of the cap domain, implying differences in the substrate preferences of the two enzymes. 3'-AMP binds in one end of a 15-A-long tunnel, with the adenine clamped between Phe23 and Tyr135, and the ribose 2'-hydroxyl interacting with Gln132. The importance of the clamp is confirmed with site-directed mutagenesis; mutation of Phe23 and Tyr135 individually to Ala increases K(m) by factors of 7 and 10, respectively. The structural data are consistent with a role for FtHAP in scavenging phosphate from small molecules present in host macrophage cells.
- Department of Chemistry, University of Missouri-Columbia, Columbia, MO 65211, USA.
Organizational Affiliation: