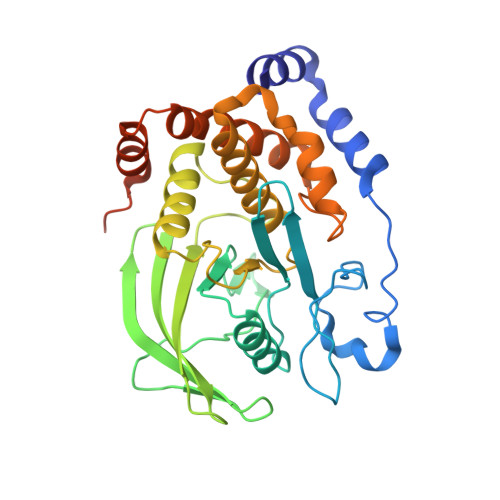
Insights into the reaction of protein-tyrosine phosphatase 1B: crystal structures for transition state analogs of both catalytic steps.
Brandao, T.A., Hengge, A.C., Johnson, S.J.(2010) J Biological Chem 285: 15874-15883
- PubMed: 20236928 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.066951
- Primary Citation Related Structures:
3I7Z, 3I80 - PubMed Abstract:
Catalysis by protein-tyrosine phosphatase 1B (PTP1B) occurs through a two-step mechanism involving a phosphocysteine intermediate. We have solved crystal structures for the transition state analogs for both steps. Together with previously reported crystal structures of apo-PTP1B, the Michaelis complex of an inactive mutant, the phosphoenzyme intermediate, and the product complex, a full picture of all catalytic steps can now be depicted. The transition state analog for the first catalytic step comprises a ternary complex between the catalytic cysteine of PTP1B, vanadate, and the peptide DADEYL, a fragment of a physiological substrate. The equatorial vanadate oxygen atoms bind to the P-loop, and the apical positions are occupied by the peptide tyrosine oxygen and by the PTP1B cysteine sulfur atom. The vanadate assumes a trigonal bipyramidal geometry in both transition state analog structures, with very similar apical O-O distances, denoting similar transition states for both phosphoryl transfer steps. Detailed interactions between the flanking peptide and the enzyme are discussed.
- Department of Chemistry and Biochemistry, Utah State University, Logan, Utah 84322-0300, USA.
Organizational Affiliation: