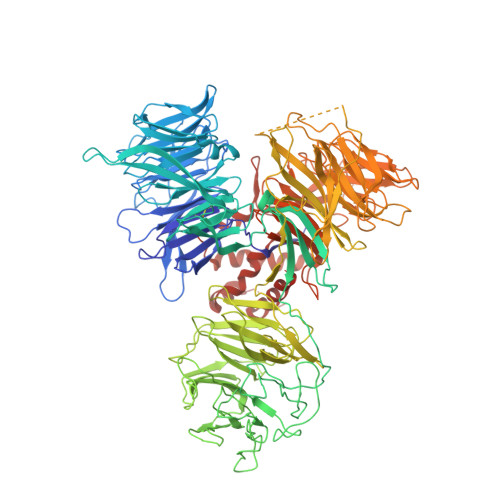
A promiscuous alpha-helical motif anchors viral hijackers and substrate receptors to the CUL4-DDB1 ubiquitin ligase machinery.
Li, T., Robert, E.I., van Breugel, P.C., Strubin, M., Zheng, N.(2010) Nat Struct Mol Biol 17: 105-111
- PubMed: 19966799 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.1719
- Primary Citation Related Structures:
3I7H, 3I7K, 3I7L, 3I7N, 3I7O, 3I7P, 3I89, 3I8C, 3I8E - PubMed Abstract:
The cullin 4-DNA-damage-binding protein 1 (CUL4-DDB1) ubiquitin ligase machinery regulates diverse cellular functions and can be subverted by pathogenic viruses. Here we report the crystal structure of DDB1 in complex with a central fragment of hepatitis B virus X protein (HBx), whose DDB1-binding activity is important for viral infection. The structure reveals that HBx binds DDB1 through an alpha-helical motif, which is also found in the unrelated paramyxovirus SV5-V protein despite their sequence divergence. Our structure-based functional analysis suggests that, like SV5-V, HBx captures DDB1 to redirect the ubiquitin ligase activity of the CUL4-DDB1 E3 ligase. We also identify the alpha-helical motif shared by these viral proteins in the cellular substrate-recruiting subunits of the E3 complex, the DDB1-CUL4-associated factors (DCAFs) that are functionally mimicked by the viral hijackers. Together, our studies reveal a common yet promiscuous structural element that is important for the assembly of cellular and virally hijacked CUL4-DDB1 E3 complexes.
- Department of Pharmacology, University of Washington, Seattle, Washington, USA.
Organizational Affiliation: