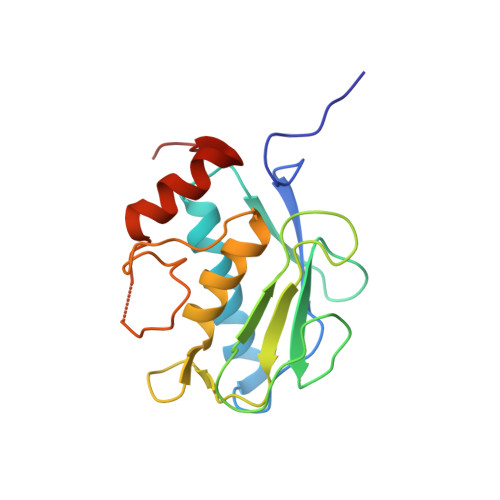
Improving potency and selectivity of a new class of non-Zn-chelating MMP-13 inhibitors.
Heim-Riether, A., Taylor, S.J., Liang, S., Gao, D.A., Xiong, Z., Michael August, E., Collins, B.K., Farmer, B.T., Haverty, K., Hill-Drzewi, M., Junker, H.D., Mariana Margarit, S., Moss, N., Neumann, T., Proudfoot, J.R., Keenan, L.S., Sekul, R., Zhang, Q., Li, J., Farrow, N.A.(2009) Bioorg Med Chem Lett 19: 5321-5324
- PubMed: 19692239 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2009.07.151
- Primary Citation Related Structures:
3I7G, 3I7I - PubMed Abstract:
Discovery and optimization of potency and selectivity of a non-Zn-chelating MMP-13 inhibitor with the aid of protein co-crystal structural information is reported. This inhibitor was observed to have a binding mode distinct from previously published MMP-13 inhibitors. Potency and selectivity were improved by extending the hit structure out from the active site into the S1' pocket.
- Boehringer Ingelheim Pharmaceuticals, Department of Medicinal Chemistry, 900 Ridgebury Rd, Ridgefield, CT 06877, USA. alexander.heim-riether@boehringer-ingelheim.com
Organizational Affiliation: