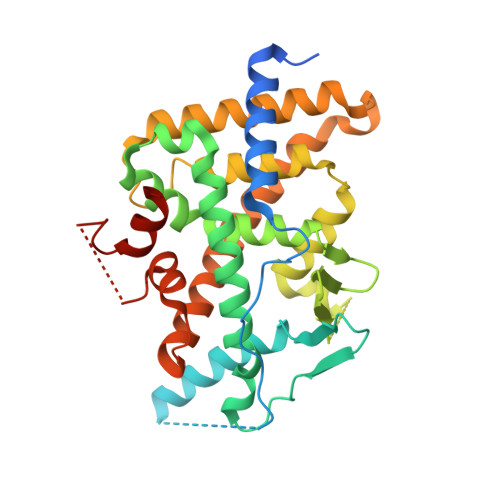
Construction and characterization of a fully active PXR/SRC-1 tethered protein with increased stability
Wang, W., Prosise, W.W., Chen, J., Taremi, S.S., Le, H.V., Madison, V., Cui, X., Thomas, A., Cheng, K.C., Lesburg, C.A.(2008) Protein Eng Des Sel 21: 425-433
- PubMed: 18456871 Search on PubMed
- DOI: https://doi.org/10.1093/protein/gzn017
- Primary Citation Related Structures:
3CTB, 3HVL - PubMed Abstract:
The nuclear xenobiotic receptor PXR is a ligand-inducible transcription factor regulating drug-metabolizing enzymes and transporters and a master switch mediating potentially adverse drug-drug interactions. In addition to binding a coactivator protein such as SRC-1, the C-terminal ligand-binding domain (LBD) is solely responsible for ligand recognition and thus the ligand-dependent downstream effects. In an effort to facilitate structural studies of PXR to understand and abolish the interactions between PXR and its ligands, several recombinant PXR/SRC-1 constructs were designed and evaluated for expression, stability and activity. Expression strategies employing either dual expression or translationally coupled bicistronic expression were found to be unsuitable for producing stable PXR in a stochiometric complex with a peptide derived from SRC-1 (SRC-1p). A single polypeptide chain encompassing PXR and SRC-1p tethered with a peptidyl linker was designed to promote intramolecular complex formation. This tethered protein was overexpressed as a soluble protein and required no additional SRC-1p for further stabilization. X-ray crystal structures in the presence and absence of the known PXR agonist SR-12813 were determined to high resolution. In addition, a circular dichroism-based binding assay was developed to allow rapid evaluation of PXR ligand affinity, making this tethered protein a convenient and effective reagent for the rational attenuation of drug-induced PXR-mediated metabolism.
- Structural Chemistry Department, Schering-Plough Research Institute, Kenilworth, NJ 07033, USA. wenyan.wang@spcorp.com
Organizational Affiliation: