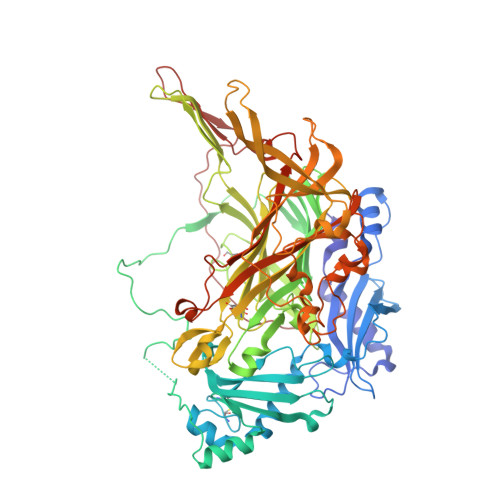
Structure and inhibition of human diamine oxidase
McGrath, A.P., Hilmer, K.M., Collyer, C.A., Shepard, E.M., Elmore, B.O., Brown, D.E., Dooley, D.M., Guss, J.M.(2009) Biochemistry 48: 9810-9822
- PubMed: 19764817 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/bi9014192
- Primary Citation Related Structures:
3HI7, 3HIG, 3HII - PubMed Abstract:
Humans have three functioning genes that encode copper-containing amine oxidases. The product of the AOC1 gene is a so-called diamine oxidase (hDAO), named for its substrate preference for diamines, particularly histamine. hDAO has been cloned and expressed in insect cells and the structure of the native enzyme determined by X-ray crystallography to a resolution of 1.8 A. The homodimeric structure has the archetypal amine oxidase fold. Two active sites, one in each subunit, are characterized by the presence of a copper ion and a topaquinone residue formed by the post-translational modification of a tyrosine. Although hDAO shares 37.9% sequence identity with another human copper amine oxidase, semicarbazide sensitive amine oxidase or vascular adhesion protein-1, its substrate binding pocket and entry channel are distinctly different in accord with the different substrate specificities. The structures of two inhibitor complexes of hDAO, berenil and pentamidine, have been refined to resolutions of 2.1 and 2.2 A, respectively. They bind noncovalently in the active-site channel. The inhibitor binding suggests that an aspartic acid residue, conserved in all diamine oxidases but absent from other amine oxidases, is responsible for the diamine specificity by interacting with the second amino group of preferred diamine substrates.
- School of Molecular and Microbial Biosciences, University of Sydney, Sydney, NSW 2006, Australia.
Organizational Affiliation: