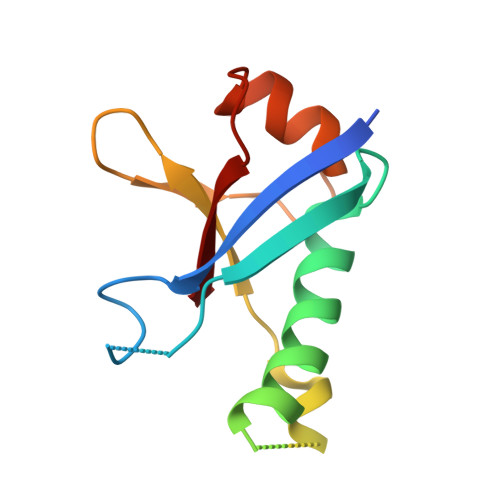
Ring1B contains a ubiquitin-like docking module for interaction with Cbx proteins.
Bezsonova, I., Walker, J.R., Bacik, J.P., Duan, S., Dhe-Paganon, S., Arrowsmith, C.H.(2009) Biochemistry 48: 10542-10548
- PubMed: 19791798 Search on PubMed
- DOI: https://doi.org/10.1021/bi901131u
- Primary Citation Related Structures:
3H8H - PubMed Abstract:
Polycomb group (PcG) proteins are a special set of repressive transcription factors involved in epigenetic modifications of chromatin. They form two functionally distinct groups of catalytically active complexes: Polycomb repressive complex 1 (PRC1) and 2 (PRC2). The PRC1 complex is an important yet poorly characterized multiprotein histone ubiquitylation machine responsible for maintaining transcriptionally silent states of genes through histone H2A K119 modification. The Ring domain containing subunits of PRC1 also have substrate-targeting domains that interact with Cbx proteins, which have been implicated in chromatin and RNA binding. In this work, we present a high resolution structure of the C-terminal domain of Ring1B, revealing a variant ubiquitin-like fold with a distinct conserved surface region. On the basis of crystal structure and mutational analysis of this domain we show that the conserved surface is responsible for interaction with Cbx members of the PRC1 and homodimer formation. These data suggest a mechanism by which Ring1B serves as an adaptor that mediates binding between the members of the PRC1 complex and the nucleosome.
- Structural Genomics Consortium, University of Toronto, 101 College Street, Toronto, Ontario, M5G 1L5, Canada.
Organizational Affiliation: