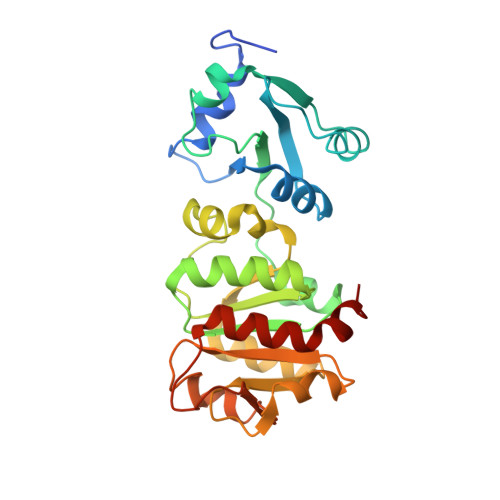
Structure of the thiostrepton-resistance methyltransferase-S-adenosyl-L-methionine complex and its interaction with ribosomal RNA
Dunstan, M.S., Hang, P.C., Zelinskaya, N.V., Honek, J.F., Conn, G.L.(2009) J Biological Chem 284: 17013-17020
- PubMed: 19369248 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M901618200
- Primary Citation Related Structures:
3GYQ - PubMed Abstract:
The x-ray crystal structure of the thiostrepton resistance RNA methyltransferase (Tsr).S-adenosyl-L-methionine (AdoMet) complex was determined at 2.45-A resolution. Tsr is definitively confirmed as a Class IV methyltransferase of the SpoU family with an N-terminal "L30-like" putative target recognition domain. The structure and our in vitro analysis of the interaction of Tsr with its target domain from 23 S ribosomal RNA (rRNA) demonstrate that the active biological unit is a Tsr homodimer. In vitro methylation assays show that Tsr activity is optimal against a 29-nucleotide hairpin rRNA though the full 58-nucleotide L11-binding domain and intact 23 S rRNA are also effective substrates. Molecular docking experiments predict that Tsr.rRNA binding is dictated entirely by the sequence and structure of the rRNA hairpin containing the A1067 target nucleotide and is most likely driven primarily by large complementary electrostatic surfaces. One L30-like domain is predicted to bind the target loop and the other is near an internal loop more distant from the target site where a nucleotide change (U1061 to A) also decreases methylation by Tsr. Furthermore, a predicted interaction with this internal loop by Tsr amino acid Phe-88 was confirmed by mutagenesis and RNA binding experiments. We therefore propose that Tsr achieves its absolute target specificity using the N-terminal domains of each monomer in combination to recognize the two distinct structural elements of the target rRNA hairpin such that both Tsr subunits contribute directly to the positioning of the target nucleotide on the enzyme.
- From the Manchester Interdisciplinary Biocentre, Faculty of Life Sciences, The University of Manchester, 131 Princess Street, Manchester M1 7DN, United Kingdom.
Organizational Affiliation: