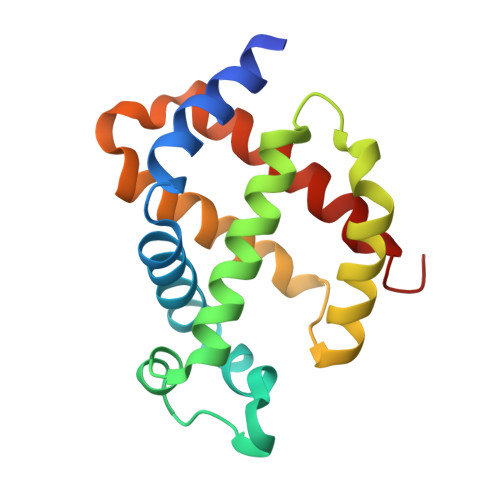
The structure of neuroglobin at high Xe and Kr pressure reveals partial conservation of globin internal cavities.
Moschetti, T., Mueller, U., Schulze, J., Brunori, M., Vallone, B.(2009) Biophys J 97: 1700-1708
- PubMed: 19751675 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.bpj.2009.05.059
- Primary Citation Related Structures:
3GK9, 3GKT, 3GLN - PubMed Abstract:
Neuroglobin (Ngb) is a hexacoordinate globin expressed in the brain of vertebrates. Ferrous Ngb binds dioxygen with high affinity and the O(2) adduct is able to scavenge NO. Convincing in vitro and in vivo data indicate that Ngb is involved in neuroprotection during hypoxia and ischemia. The 3D structure of Ngb reveals the presence of a wide internal cavity connecting its heme active site with the bulk. To explore the role of this "tunnel" in the control of ligand binding, we determined the structure of metNgb and NgbCO equilibrated with Xe or Kr. We show four docking sites for Xe (only two for Kr); two of the four Xe sites are within the large cavity. They are only partially conserved in globins, since the two proximal Xe sites identified in myoglobin (Xe1 and Xe2) are absent in Ngb, as well as in cytoglobin. The Xe docking sites in Ngb map a pathway within the protein matrix, leading to the heme, which becomes more accessible in the ligand-bound species. This may be of significance in connection with the redox chemistry that may be the primary function of this hexacoordinate globin.
- Department of Biochemical Sciences A.Rossi-Fanelli, University of Rome La Sapienza, Rome, Italy.
Organizational Affiliation: