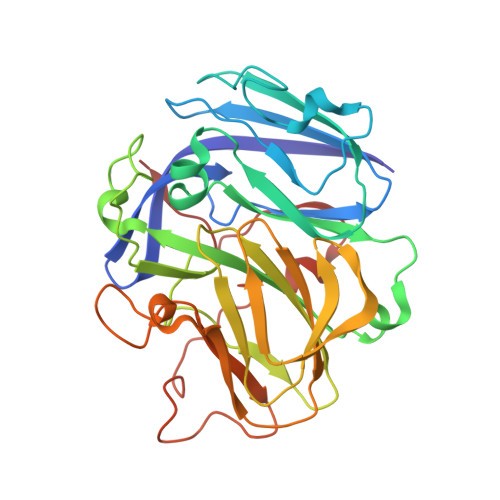
Crystal structure of a two-domain multicopper oxidase: implications for the evolution of multicopper blue proteins.
Lawton, T.J., Sayavedra-Soto, L.A., Arp, D.J., Rosenzweig, A.C.(2009) J Biological Chem 284: 10174-10180
- PubMed: 19224923 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M900179200
- Primary Citation Related Structures:
3G5W - PubMed Abstract:
The two-domain multicopper oxidases are proposed to be key intermediates in the evolution of three-domain multicopper oxidases. A number of two-domain multicopper oxidases have been identified from genome sequences and are classified as type A, type B, or type C on the basis of the predicted location of the type 1 copper center. The crystal structure of blue copper oxidase, a type C two-domain multicopper oxidase from Nitrosomonas europaea, has been determined to 1.9 A resolution. Blue copper oxidase is a trimer, of which each subunit comprises two cupredoxin domains. Each subunit houses a type 1 copper site in domain 1 and a type 2/type 3 trinuclear copper cluster at the subunit-subunit interface. The coordination geometry at the trinuclear copper site is consistent with reduction of the copper ions. Although the overall architecture of blue copper oxidase is similar to nitrite reductases, detailed structural alignments show that the fold and domain orientation more closely resemble the three-domain multicopper oxidases. These observations have important implications for the evolution of nitrite reductases and multicopper oxidases.
- Departments of Biochemistry, Molecular Biology, and Cell Biology and of Chemistry, Northwestern University, Evanston, Illinois 60208, USA.
Organizational Affiliation: