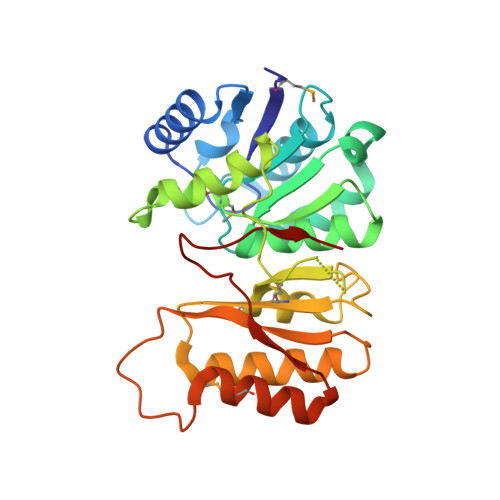
Structure and function of a novel LD-carboxypeptidase a involved in peptidoglycan recycling.
Das, D., Herve, M., Elsliger, M.A., Kadam, R.U., Grant, J.C., Chiu, H.J., Knuth, M.W., Klock, H.E., Miller, M.D., Godzik, A., Lesley, S.A., Deacon, A.M., Mengin-Lecreulx, D., Wilson, I.A.(2013) J Bacteriol 195: 5555-5566
- PubMed: 24123814 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00900-13
- Primary Citation Related Structures:
3G23 - PubMed Abstract:
Approximately 50% of cell wall peptidoglycan in Gram-negative bacteria is recycled with each generation. The primary substrates used for peptidoglycan biosynthesis and recycling in the cytoplasm are GlcNAc-MurNAc(anhydro)-tetrapeptide and its degradation product, the free tetrapeptide. This complex process involves ∼15 proteins, among which the cytoplasmic enzyme ld-carboxypeptidase A (LdcA) catabolizes the bond between the last two l- and d-amino acid residues in the tetrapeptide to form the tripeptide, which is then utilized as a substrate by murein peptide ligase (Mpl). LdcA has been proposed as an antibacterial target. The crystal structure of Novosphingobium aromaticivorans DSM 12444 LdcA (NaLdcA) was determined at 1.89-Å resolution. The enzyme was biochemically characterized and its interactions with the substrate modeled, identifying residues potentially involved in substrate binding. Unaccounted electron density at the dimer interface in the crystal suggested a potential site for disrupting protein-protein interactions should a dimer be required to perform its function in bacteria. Our analysis extends the identification of functional residues to several other homologs, which include enzymes from bacteria that are involved in hydrocarbon degradation and destruction of coral reefs. The NaLdcA crystal structure provides an alternate system for investigating the structure-function relationships of LdcA and increases the structural coverage of the protagonists in bacterial cell wall recycling.
- Joint Center for Structural Genomics‡
Organizational Affiliation: