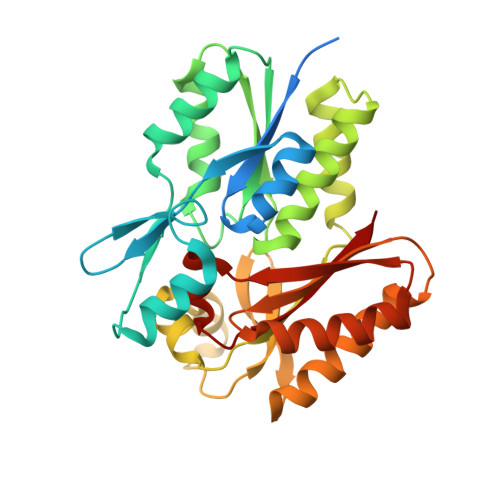
Structure of a fatty acid-binding protein from Bacillus subtilis determined by sulfur-SAD phasing using in-house chromium radiation
Nan, J., Zhou, Y.F., Yang, C., Brostromer, E., Kristensen, O., Su, X.-D.(2009) Acta Crystallogr D Biol Crystallogr 65: 440-448
- PubMed: 19390149 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444909007756
- Primary Citation Related Structures:
3FYS - PubMed Abstract:
Sulfur single-wavelength anomalous dispersion (S-SAD) and halide-soaking methods are increasingly being used for ab initio phasing. With the introduction of in-house Cr X-ray sources, these methods benefit from the enhanced anomalous scattering of S and halide atoms, respectively. Here, these methods were combined to determine the crystal structure of BsDegV, a DegV protein-family member from Bacillus subtilis. The protein was cocrystallized with bromide and low-redundancy data were collected to 2.5 A resolution using Cr Kalpha radiation. 17 heavy-atom sites (ten sulfurs and seven bromides) were located using standard methods. The anomalous scattering of some of the BsDegV S atoms and Br atoms was weak, thus neither sulfurs nor bromides could be used alone for structure determination using the collected data. When all 17 heavy-atom sites were used for SAD phasing, an easily interpretable electron-density map was obtained after density modification. The model of BsDegV was built automatically and a palmitate was found tightly bound in the active site. Sequence alignment and comparisons with other known DegV structures provided further insight into the specificity of fatty-acid selection and recognition within this protein family.
- National Laboratory of Protein Engineering and Plant Genetic Engineering, College of Life Sciences, Peking University, Beijing, People's Republic of China.
Organizational Affiliation: