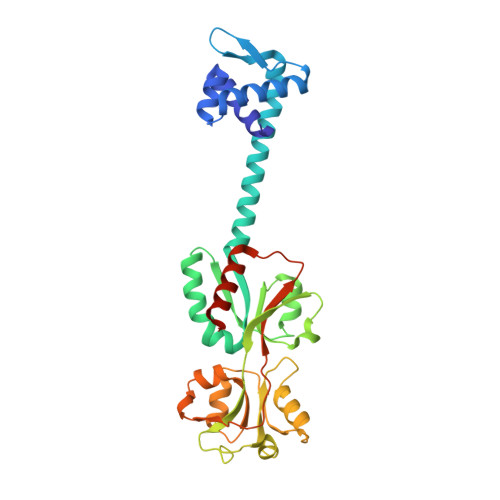
Structural studies on the full-length LysR-type regulator TsaR from Comamonas testosteroni T-2 reveal a novel open conformation of the tetrameric LTTR fold
Monferrer, D., Tralau, T., Kertesz, M.A., Dix, I., Sola, M., Uson, I.(2010) Mol Microbiol 75: 1199-1214
- PubMed: 20059681 Search on PubMed
- DOI: https://doi.org/10.1111/j.1365-2958.2010.07043.x
- Primary Citation Related Structures:
3FXQ, 3FXR, 3FXU, 3FZJ - PubMed Abstract:
LysR-type transcriptional regulators (LTTRs) constitute the largest family of regulators in prokaryotes. The full-length structures of the LTTR TsaR from Comamonas testosteroni T-2 and its complex with the natural inducer para-toluensulfonate have been characterized by X-ray diffraction. Both ligand-free and complexed forms reveal a dramatically different quaternary structure from that of CbnR from Ralstonia eutropha, or a putative LysR-type regulator from Pseudomonas aeruginosa, the only other determined full-length structures of tetrameric LTTRs. Although all three show a head-to-head tetrameric ring, TsaR displays an open conformation, whereas CbnR and PA01-PR present additional contacts in opposing C-terminal domains that close the ring. Such large differences may be due to a broader structural versatility than previously assumed or either, reflect the intrinsic flexibility of tetrameric LTTRs. On the grounds of the sliding dimer hypothesis of LTTR activation, we propose a structural model in which the closed structures could reflect the conformation of a ligand-free LTTR, whereas inducer binding would bring about local changes to disrupt the interface linking the two compact C-terminal domains. This could lead to a TsaR-like, open structure, where the pairs of recognition helices are closer to each other by more than 10 A.
- IBMB-CSIC, Baldiri Reixach 15, Barcelona Science Park, 08028, Barcelona, Spain.
Organizational Affiliation: