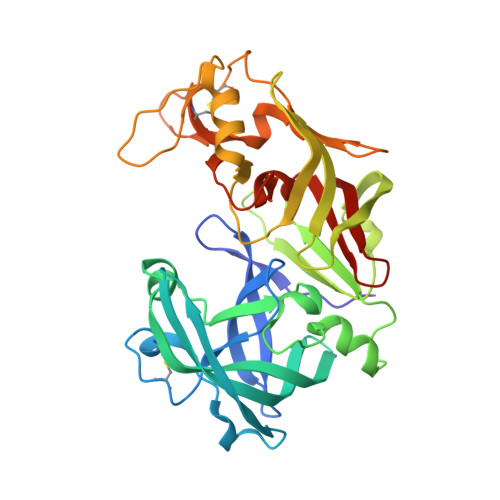
Crystal structures of the histo-aspartic protease (HAP) from Plasmodium falciparum.
Bhaumik, P., Xiao, H., Parr, C.L., Kiso, Y., Gustchina, A., Yada, R.Y., Wlodawer, A.(2009) J Mol Biology 388: 520-540
- PubMed: 19285084 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2009.03.011
- Primary Citation Related Structures:
3FNS, 3FNT, 3FNU - PubMed Abstract:
The structures of recombinant histo-aspartic protease (HAP) from malaria-causing parasite Plasmodium falciparum as apoenzyme and in complex with two inhibitors, pepstatin A and KNI-10006, were solved at 2.5-, 3.3-, and 3.05-A resolutions, respectively. In the apoenzyme crystals, HAP forms a tight dimer not seen previously in any aspartic protease. The interactions between the monomers affect the conformation of two flexible loops, the functionally important "flap" (residues 70-83) and its structural equivalent in the C-terminal domain (residues 238-245), as well as the orientation of helix 225-235. The flap is found in an open conformation in the apoenzyme. Unexpectedly, the active site of the apoenzyme contains a zinc ion tightly bound to His32 and Asp215 from one monomer and to Glu278A from the other monomer, with the coordination of Zn resembling that seen in metalloproteases. The flap is closed in the structure of the pepstatin A complex, whereas it is open in the complex with KNI-10006. Although the binding mode of pepstatin A is significantly different from that in other pepsin-like aspartic proteases, its location in the active site makes unlikely the previously proposed hypothesis that HAP is a serine protease. The binding mode of KNI-10006 is unusual compared with the binding of other inhibitors from the KNI series to aspartic proteases. The novel features of the HAP active site could facilitate design of specific inhibitors used in the development of antimalarial drugs.
- Protein Structure Section, Macromolecular Crystallography Laboratory, National Cancer Institute, Frederick, MD 21702, USA.
Organizational Affiliation: