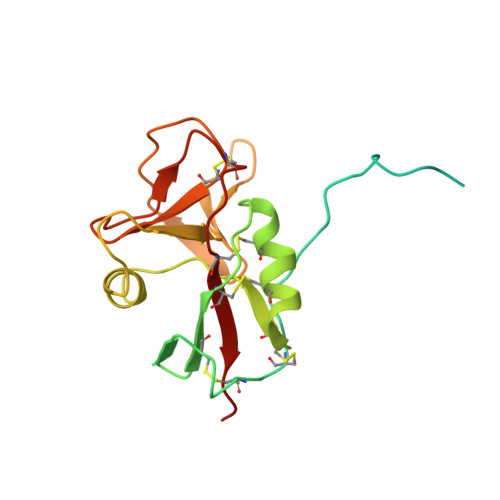
Structure of Epstein-Barr virus glycoprotein 42 suggests a mechanism for triggering receptor-activated virus entry.
Kirschner, A.N., Sorem, J., Longnecker, R., Jardetzky, T.S.(2009) Structure 17: 223-233
- PubMed: 19217393 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2008.12.010
- Primary Citation Related Structures:
3FD4 - PubMed Abstract:
Epstein-Barr virus requires glycoproteins gH/gL, gB, and gp42 to fuse its lipid envelope with B cells. Gp42 is a type II membrane protein consisting of a flexible N-terminal region, which binds gH/gL, and a C-terminal lectin-like domain that binds to the B-cell entry receptor human leukocyte antigen (HLA) class II. Gp42 triggers membrane fusion after HLA binding, a process that requires simultaneous binding to gH/gL and a functional hydrophobic pocket in the lectin domain adjacent to the HLA binding site. Here we present the structure of gp42 in its unbound form. Comparisons to the previously determined structure of a gp42:HLA complex reveals additional N-terminal residues forming part of the gH/gL binding site and structural changes in the receptor binding domain. Although the core of the lectin domain remains similar, significant shifts in two loops and an alpha helix bordering the essential hydrophobic pocket suggest a structural mechanism for triggering fusion.
- Department of Biochemistry, Molecular Biology, and Cell Biology, Northwestern University, Evanston, IL 60208, USA.
Organizational Affiliation: