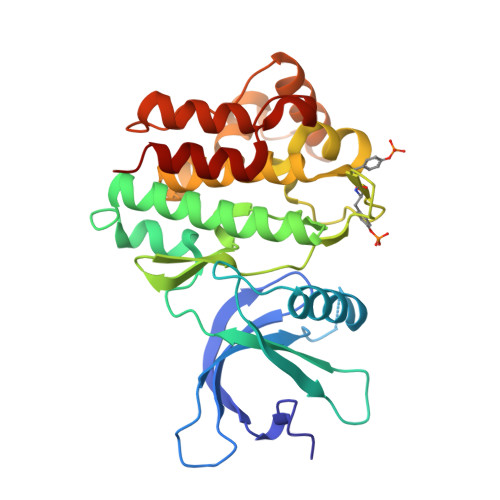
Dissecting specificity in the Janus kinases: the structures of JAK-specific inhibitors complexed to the JAK1 and JAK2 protein tyrosine kinase domains.
Williams, N.K., Bamert, R.S., Patel, O., Wang, C., Walden, P.M., Wilks, A.F., Fantino, E., Rossjohn, J., Lucet, I.S.(2009) J Mol Biology 387: 219-232
- PubMed: 19361440 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2009.01.041
- Primary Citation Related Structures:
3EYG, 3EYH, 3FUP - PubMed Abstract:
The Janus kinases (JAKs) are a pivotal family of protein tyrosine kinases (PTKs) that play prominent roles in numerous cytokine signaling pathways, with aberrant JAK activity associated with a variety of hematopoietic malignancies, cardiovascular diseases and immune-related disorders. Whereas the structures of the JAK2 and JAK3 PTK domains have been determined, the structure of the JAK1 PTK domain is unknown. Here, we report the high-resolution crystal structures of the "active form" of the JAK1 PTK domain in complex with two JAK inhibitors, a tetracyclic pyridone 2-t-butyl-9-fluoro-3,6-dihydro-7H-benz[h]-imidaz[4,5-f]isoquinoline-7-one (CMP6) and (3R,4R)-3-[4-methyl-3-[N-methyl-N-(7H-pyrrolo[2,3-d]pyrimidin-4-yl)amino]piperidin-1-yl]-3-oxopropionitrile (CP-690,550), and compare them with the corresponding JAK2 PTK inhibitor complexes. Both inhibitors bound in a similar manner to JAK1, namely buried deep within a constricted ATP-binding site, thereby providing a basis for the potent inhibition of JAK1. As expected, the mode of inhibitor binding in JAK1 was very similar to that observed in JAK2, highlighting the challenges in developing JAK-specific inhibitors that target the ATP-binding site. Nevertheless, differences surrounding the JAK1 and JAK2 ATP-binding sites were apparent, thereby providing a platform for the rational design of JAK2- and JAK1-specific inhibitors.
- Protein Crystallography Unit, Department of Biochemistry and Molecular Biology, School of Biomedical Sciences, Monash University, Clayton, Victoria, Australia.
Organizational Affiliation: