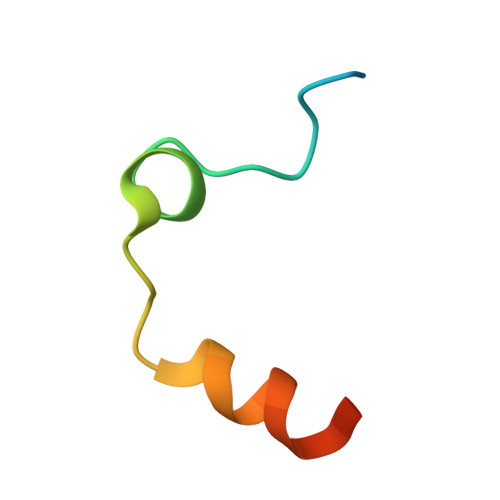
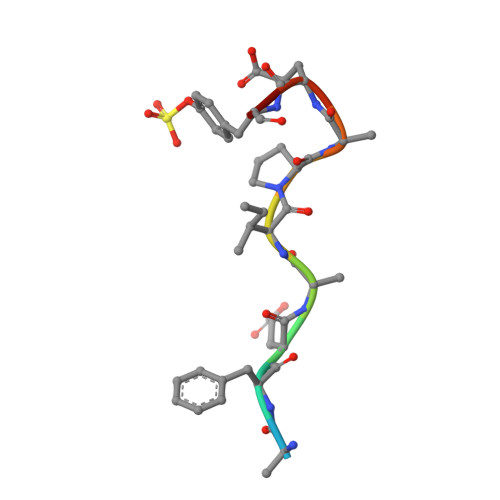
KNOBLE: a knowledge-based approach for the design and synthesis of readily accessible small-molecule chemical probes to test protein binding
Gerlach, C., Munzel, M., Baum, B., Gerber, H.-D., Craan, T., Diederich, W.E., Klebe, G.(2007) Angew Chem Int Ed Engl 46: 9105-9109
- PubMed: 17955562 Search on PubMed
- DOI: https://doi.org/10.1002/anie.200703323
- Primary Citation Related Structures:
3EGK - Institut für Pharmazeutische Chemie, Philipps-Universität Marburg, Marbacher Weg 6, 35032 Marburg, Germany.
Organizational Affiliation: