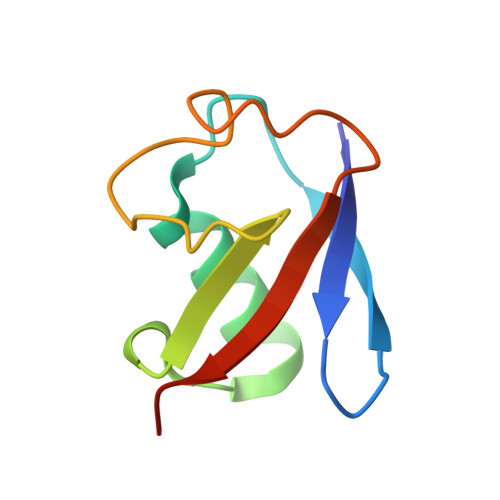
Structural probing of Zn(II), Cd(II) and Hg(II) binding to human ubiquitin.
Falini, G., Fermani, S., Tosi, G., Arnesano, F., Natile, G.(2008) Chem Commun (Camb) 45: 5960-5962
- PubMed: 19030552
- DOI: https://doi.org/10.1039/b813463d
- Primary Citation Related Structures:
3EEC, 3EFU, 3EHV - PubMed Abstract:
A structural investigation performed on adducts of human ubiquitin with group-12 metal ions reveals common preferential anchoring sites, the most populated one being His68; at higher metal ion concentration a second and a third site, close to the N-terminus of the protein, become populated and promote a polymorphic transition from orthorhombic to cubic form; Glu16 and Glu18, involved in the latter metal binding, undergo a remarkable displacement from their position in native ubiquitin; the aggregate stereochemistry appears to be driven by the clustering of deshielded backbone hydrogen-bond patches, and metal ions foster this process.
- Dipartimento di Chimica, G. Ciamician, Università di Bologna, via Selmi 2, 40126, Bologna, Italy. giuseppe.falini@unibo.it
Organizational Affiliation: