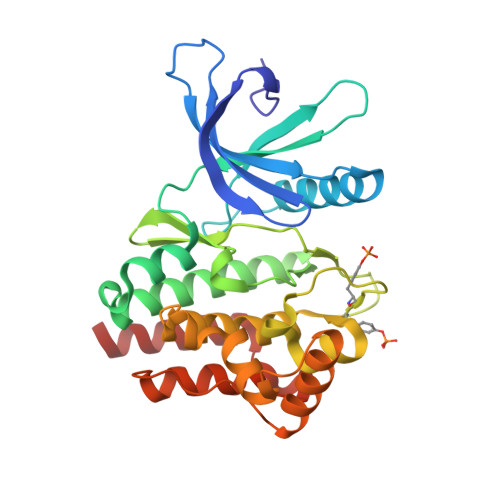
Fragment-based discovery of JAK-2 inhibitors.
Antonysamy, S., Hirst, G., Park, F., Sprengeler, P., Stappenbeck, F., Steensma, R., Wilson, M., Wong, M.(2009) Bioorg Med Chem Lett 19: 279-282
- PubMed: 19019674 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2008.08.064
- Primary Citation Related Structures:
3E62, 3E63, 3E64 - PubMed Abstract:
Fragment-based hit identification coupled with crystallographically enabled structure-based drug design was used to design potent inhibitors of JAK-2. After two iterations from fragment 1, we were able to increase potency by greater than 500-fold to provide sulfonamide 13, a 78-nM JAK-2 inhibitor.
- Medicinal Chemistry, SGX Pharmaceuticals, Inc, San Diego, CA 92121, USA.
Organizational Affiliation: