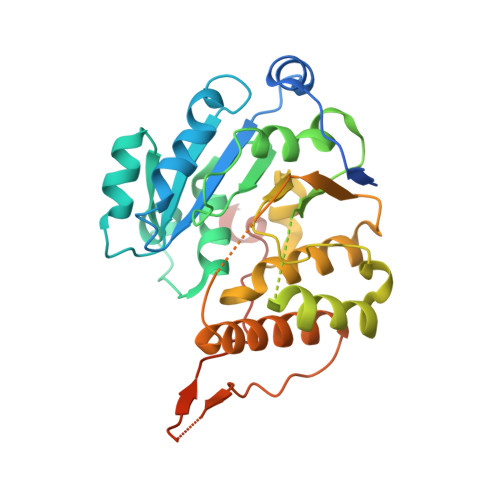
Mycobacterium tuberculosis glucosyl-3-phosphoglycerate synthase: structure of a key enzyme in methylglucose lipopolysaccharide biosynthesis
Pereira, P.J.B., Empadinhas, N., Albuquerque, L., Sa-Moura, B., da Costa, M.S., Macedo-Ribeiro, S.(2008) PLoS One 3: e3748-e3748
- PubMed: 19015727 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pone.0003748
- Primary Citation Related Structures:
3E25, 3E26 - PubMed Abstract:
Tuberculosis constitutes today a serious threat to human health worldwide, aggravated by the increasing number of identified multi-resistant strains of Mycobacterium tuberculosis, its causative agent, as well as by the lack of development of novel mycobactericidal compounds for the last few decades. The increased resilience of this pathogen is due, to a great extent, to its complex, polysaccharide-rich, and unusually impermeable cell wall. The synthesis of this essential structure is still poorly understood despite the fact that enzymes involved in glycosidic bond synthesis represent more than 1% of all M. tuberculosis ORFs identified to date. One of them is GpgS, a retaining glycosyltransferase (GT) with low sequence homology to any other GTs of known structure, which has been identified in two species of mycobacteria and shown to be essential for the survival of M. tuberculosis. To further understand the biochemical properties of M. tuberculosis GpgS, we determined the three-dimensional structure of the apo enzyme, as well as of its ternary complex with UDP and 3-phosphoglycerate, by X-ray crystallography, to a resolution of 2.5 and 2.7 A, respectively. GpgS, the first enzyme from the newly established GT-81 family to be structurally characterized, displays a dimeric architecture with an overall fold similar to that of other GT-A-type glycosyltransferases. These three-dimensional structures provide a molecular explanation for the enzyme's preference for UDP-containing donor substrates, as well as for its glucose versus mannose discrimination, and uncover the structural determinants for acceptor substrate selectivity. Glycosyltransferases constitute a growing family of enzymes for which structural and mechanistic data urges. The three-dimensional structures of M. tuberculosis GpgS now determined provide such data for a novel enzyme family, clearly establishing the molecular determinants for substrate recognition and catalysis, while providing an experimental scaffold for the structure-based rational design of specific inhibitors, which lay the foundation for the development of novel anti-tuberculosis therapies.
- Instituto de Biologia Molecular e Celular (IBMC), Universidade do Porto, Porto, Portugal.
Organizational Affiliation: