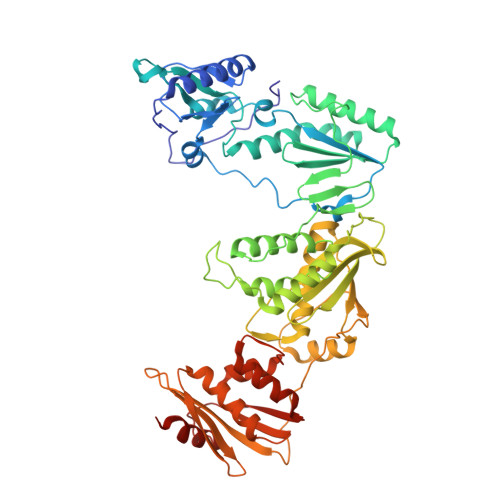
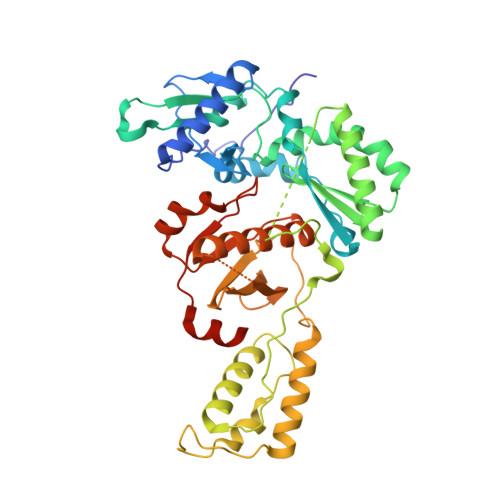
Design of annulated pyrazoles as inhibitors of HIV-1 reverse transcriptase
Sweeney, Z.K., Harris, S.F., Arora, S.F., Javanbakht, H., Li, Y., Fretland, J., Davidson, J.P., Billedeau, J.R., Gleason, S.K., Hirschfeld, D., Kennedy-Smith, J.J., Mirzadegan, T., Roetz, R., Smith, M., Sperry, S., Suh, J.M., Wu, J., Tsing, S., Villasenor, A.G., Paul, A., Su, G., Heilek, G., Hang, J.Q., Zhou, A.S., Jernelius, J.A., Zhang, F.J., Klumpp, K.(2008) J Med Chem 51: 7449-7458
- PubMed: 19007201 Search on PubMed
- DOI: https://doi.org/10.1021/jm800527x
- Primary Citation Related Structures:
3DYA, 3E01 - PubMed Abstract:
Non-nucleoside reverse transcriptase inhibitors (NNRTIs) are recommended components of preferred combination antiretroviral therapies used for the treatment of HIV. These regimens are extremely effective in suppressing virus replication. Structure-based optimization of diaryl ether inhibitors led to the discovery of a new series of pyrazolo[3,4-c]pyridazine NNRTIs that bind the reverse transcriptase enzyme of human immunodeficiency virus-1 (HIV-RT) in an expanded volume relative to most other inhibitors in this class.The binding mode maintains the beta13 and beta14 strands bearing Pro236 in a position similar to that in the unliganded reverse transcriptase structure, and the distribution of interactions creates the opportunity for substantial resilience to single point mutations. Several pyrazolopyridazine NNRTIs were found to be highly effective against wild-type and NNRTI-resistant viral strains in cell culture.
- Department of Medicinal Chemistry, Roche Palo Alto LLC, 3431 Hillview Avenue, Palo Alto, California 94304, USA. zachary.sweeney@roche.com
Organizational Affiliation: