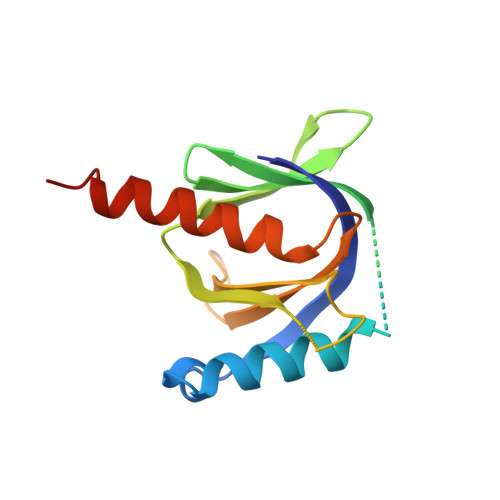
Crystal structure of the human Fe65-PTB1 domain.
Radzimanowski, J., Ravaud, S., Schlesinger, S., Koch, J., Beyreuther, K., Sinning, I., Wild, K.(2008) J Biological Chem 283: 23113-23120
- PubMed: 18550529 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M800861200
- Primary Citation Related Structures:
3D8D, 3D8E, 3D8F - PubMed Abstract:
The neuronal adaptor protein Fe65 is involved in brain development, Alzheimer disease amyloid precursor protein (APP) signaling, and proteolytic processing of APP. It contains three protein-protein interaction domains, one WW domain, and a unique tandem array of phosphotyrosine-binding (PTB) domains. The N-terminal PTB domain (Fe65-PTB1) was shown to interact with a variety of proteins, including the low density lipoprotein receptor-related protein (LRP-1), the ApoEr2 receptor, and the histone acetyltransferase Tip60. We have determined the crystal structures of human Fe65-PTB1 in its apo- and in a phosphate-bound form at 2.2 and 2.7A resolution, respectively. The overall fold shows a PTB-typical pleckstrin homology domain superfold. Although Fe65-PTB1 has been classified on an evolutionary basis as a Dab-like PTB domain, it contains attributes of other PTB domain subfamilies. The phosphotyrosine-binding pocket resembles IRS-like PTB domains, and the bound phosphate occupies the binding site of the phosphotyrosine (Tyr(P)) within the canonical NPXpY recognition motif. In addition Fe65-PTB1 contains a loop insertion between helix alpha2 and strand beta2(alpha2/beta2 loop) similar to members of the Shc-like PTB domain subfamily. The structural comparison with the Dab1-PTB domain reveals a putative phospholipid-binding site opposite the peptide binding pocket. We suggest Fe65-PTB1 to interact with its target proteins involved in translocation and signaling of APP in a phosphorylation-dependent manner.
- Heidelberg University Biochemistry Center, INF328, D-69120 Heidelberg, Germany.
Organizational Affiliation: