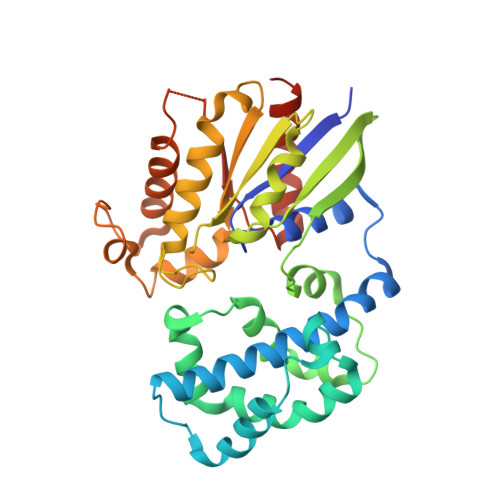
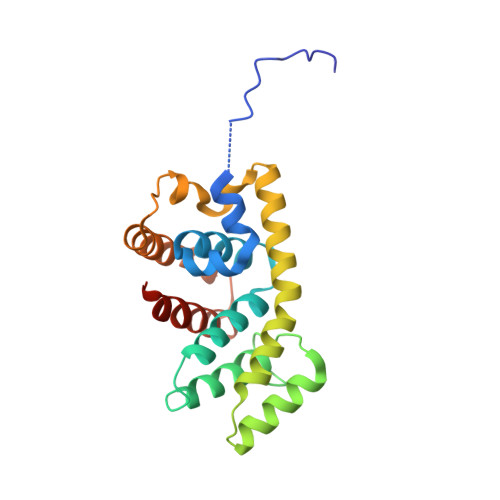
Recognition of the Activated States of Galpha13 by the rgRGS Domain of PDZRhoGEF.
Chen, Z., Singer, W.D., Danesh, S.M., Sternweis, P.C., Sprang, S.R.(2008) Structure 16: 1532-1543
- PubMed: 18940608 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2008.07.009
- Primary Citation Related Structures:
3CX6, 3CX7, 3CX8 - PubMed Abstract:
G12 class heterotrimeric G proteins stimulate RhoA activation by RGS-RhoGEFs. However, p115RhoGEF is a GTPase Activating Protein (GAP) toward Galpha13, whereas PDZRhoGEF is not. We have characterized the interaction between the PDZRhoGEF rgRGS domain (PRG-rgRGS) and the alpha subunit of G13 and have determined crystal structures of their complexes in both the inactive state bound to GDP and the active states bound to GDP*AlF (transition state) and GTPgammaS (Michaelis complex). PRG-rgRGS interacts extensively with the helical domain and the effector-binding sites on Galpha13 through contacts that are largely conserved in all three nucleotide-bound states, although PRG-rgRGS has highest affinity to the Michaelis complex. An acidic motif in the N terminus of PRG-rgRGS occupies the GAP binding site of Galpha13 and is flexible in the GDP*AlF complex but well ordered in the GTPgammaS complex. Replacement of key residues in this motif with their counterparts in p115RhoGEF confers GAP activity.
- Department of Pharmacology, The University of Texas Southwestern Medical Center, Dallas, TX 75390, USA.
Organizational Affiliation: