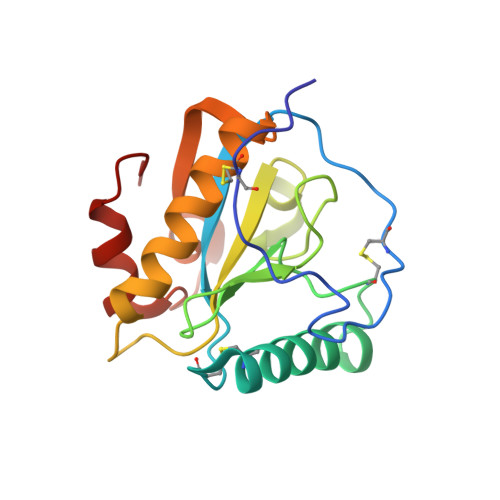
Crystal structure of the peptidoglycan recognition protein at 1.8 A resolution reveals dual strategy to combat infection through two independent functional homodimers
Sharma, P., Singh, N., Sinha, M., Sharma, S., Perbandt, M., Betzel, C., Kaur, P., Srinivasan, A., Singh, T.P.(2008) J Mol Biology 378: 921-930
- PubMed: 18395744 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.03.018
- Primary Citation Related Structures:
3C2X - PubMed Abstract:
The mammalian peptidoglycan recognition protein-S (PGRP-S) binds to peptidoglycans (PGNs), which are essential components of the cell wall of bacteria. The protein was isolated from the samples of milk obtained from camels with mastitis and purified to homogeneity and crystallized. The crystals belong to orthorhombic space group I222 with a=87.0 A, b=101.7 A and c=162.3 A having four crystallographically independent molecules in the asymmetric unit. The structure has been determined using X-ray crystallographic data and refined to 1.8 A resolution. Overall, the structures of all the four crystallographically independent molecules are identical. The folding of PGRP-S consists of a central beta-sheet with five beta-strands, four parallel and one antiparallel, and three alpha-helices. This protein fold provides two functional sites. The first of these is the PGN-binding site, located on the groove that opens on the surface in the direction opposite to the location of the N terminus. The second site is implicated to be involved in the binding of non-PGN molecules, it also includes putative N-terminal segment residues (1-31) and helix alpha2 in the extended binding. The structure reveals a novel arrangement of PGRP-S molecules in which two pairs of molecules associate to form two independent dimers. The first dimer is formed by two molecules with N-terminal segments at the interface in which non-PGN binding sites are buried completely, whereas the PGN-binding sites of two participating molecules are fully exposed at the opposite ends of the dimer. In the second dimer, PGN-binding sites are buried at the interface while non-PGN binding sites are fully exposed at the opposite ends of the dimer. This form of dimeric arrangement is unique and seems to be aimed at enhancing the capability of the protein against specific invading bacteria. This mode of functional dimerization enhances efficiency and specificity, and is observed for the first time in the family of PGRP molecules.
- Department of Biophysics, All India Institute of Medical Sciences, New Delhi, India.
Organizational Affiliation: