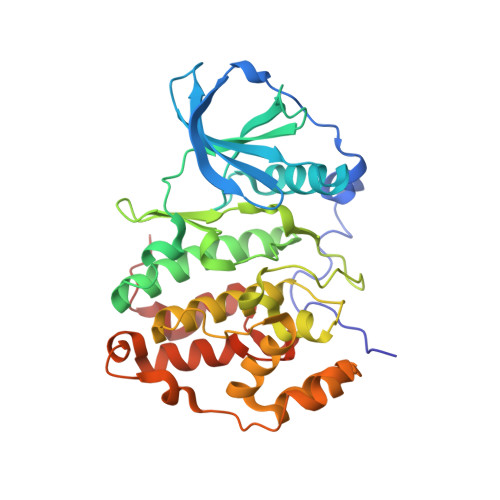
The Catalytic Subunit of Human Protein Kinase CK2 Structurally Deviates from Its Maize Homologue in Complex with the Nucleotide Competitive Inhibitor Emodin
Raaf, J., Klopffleisch, K., Issinger, O.-G., Niefind, K.(2008) J Mol Biology 377: 1-8
- PubMed: 18242640 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2008.01.008
- Primary Citation Related Structures:
3BQC, 3C13 - PubMed Abstract:
The Ser/Thr kinase CK2 (former name: casein kinase 2) is a heterotetrameric enzyme composed of two catalytic chains (CK2alpha) attached to a dimer of noncatalytic subunits. Together with the cyclin-dependent kinases and the mitogen-activated protein kinases, CK2alpha belongs to the CMGC family of the eukaryotic protein kinases. CK2 is an important survival and stability factor in eukaryotic cells: its catalytic activity is elevated in a wide variety of tumors while its down-regulation can lead to apoptosis. Thus, CK2 is a valuable target for drug development and for chemical biology approaches of cell biological research, and small organic inhibitors addressing CK2 are of considerable interest. We describe here the complex structure between a C-terminal deletion mutant of human CK2alpha and the ATP-competitive inhibitor emodin (1,3,8-trihydroxy-6-methylanthraquinone, International Union of Pure and Applied Chemistry name: 1,3,8-trihydroxy-6-methylanthracene-9,10-dione) and compare it with a previously published complex structure of emodin and maize CK2alpha. With a resolution of 1.5 A, the human CK2alpha/emodin structure has a much better resolution than its maize counterpart (2.6 A). Even more important, in spite of a sequence identity of more than 77% between human and maize CK2alpha, the two structures deviate significantly in the orientation, in which emodin is trapped by the enzyme, and in the local conformations around the ligand binding site: maize CK2alpha shows its largest adaptations in the ATP-binding loop, whereas human CK2alpha shows its largest adaptations in the hinge region connecting the two main domains of the protein kinase core. These observations emphasize the importance of local plasticity for ligand binding and demonstrate that two orthologues of an enzyme can behave quite different in this respect.
- Universität zu Köln, Institut für Biochemie, Zülpicher Strasse 47, D-50674 Köln, Germany.
Organizational Affiliation: