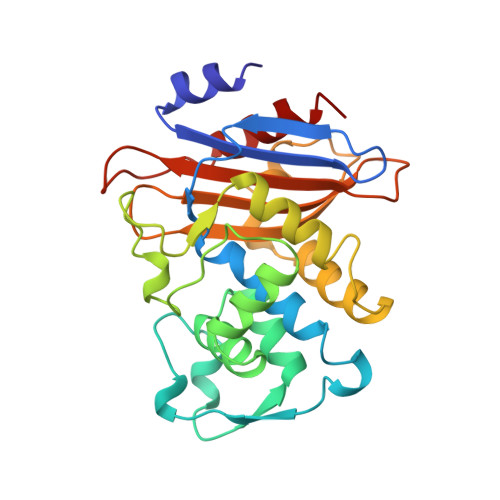
Refined crystal structure of beta-lactamase from Staphylococcus aureus PC1 at 2.0 A resolution.
Herzberg, O.(1991) J Mol Biology 217: 701-719
- PubMed: 2005620
- DOI: https://doi.org/10.1016/0022-2836(91)90527-d
- Primary Citation of Related Structures:
3BLM - PubMed Abstract:
The crystal structure of a class A beta-lactamase from Staphylococcus aureus PC1 has been refined at 2.0 A resolution. The resulting crystallographic R-factor (R = sigma h parallel Fo[-]Fc parallel/sigma h[Fo], where [Fo] and [Fc] are the observed and calculated structure factor amplitudes, respectively), is 0.163 for the 17,547 reflections with I greater than or equal to 2 sigma (I) within the 8.0 A to 2.0 A resolution range. The molecule consists of two closely associated domains. One domain is formed by a five-stranded antiparallel beta-sheet with three helices packing against a face of the sheet. The second domain is formed mostly by helices that pack against the second face of the sheet. The active site is located in the interface between the two domains, and many of the residues that form it are conserved in all known sequences of class A beta-lactamases. Similar to the serine proteases, an oxyanion hole is implicated in catalysis. It is formed by two main-chain nitrogen atoms, that of the catalytic seryl residue, Ser70, and that of Gln237 on an edge beta-strand of the major beta-sheet. Ser70 is interacting with another conserved seryl residue, Ser130, located between the two ammonium groups of the functionally important lysine residues, Lys73 and Lys234. Such intricate interactions point to a possible catalytic role for this second seryl residue. Another key catalytic residue is Glu166. There are several unusual structural features associated with the active site. (1) A cis peptide bond has been identified between the catalytic Glu166 and Ile167. (2) Ala69 and Leu220 have strained phi, psi dihedral angles making close contacts that restrict the conformation of the active site beta-strand involved in the formation of the oxyanion hole. (3) A buried aspartate residue, the conserved Asp233, is located next to the active site Lys234. It is interacting with another buried aspartyl residue, Asp246. An internal solvent molecule is also involved, but the rest of its interactions with the protein indicate it is not a cation. (4) Another conserved aspartyl residue that is desolvated is Asp131, adjacent to Ser130. Its charge is stabilized by interactions with four main-chain nitrogen atoms. (5) An internal cavity underneath the active site depression is filled with six solvent molecules. This, and an adjacent cavity occupied by three solvent molecules partially separate the omega-loop associated with the active site from the rest of the protein.(ABSTRACT TRUNCATED AT 250 WORDS)
- Center for Advanced Research in Biotechnology, Maryland Biotechnology Institute, University of Maryland, Rockville 20850.
Organizational Affiliation: