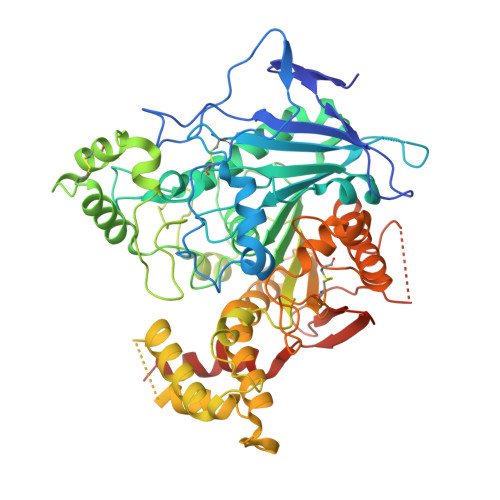
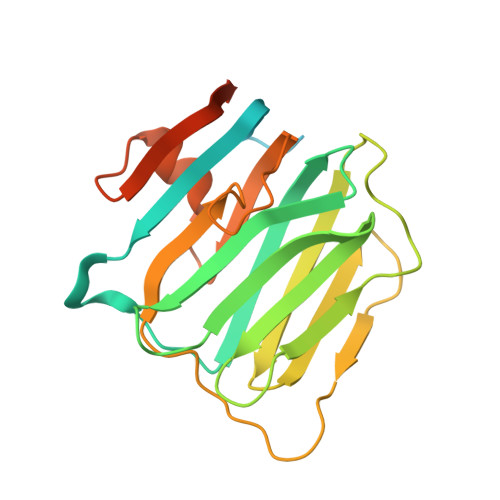
Structures of Neuroligin-1 and the Neuroligin-1/Neurexin-1beta Complex Reveal Specific Protein-Protein and Protein-Ca(2+) Interactions.
Arac, D., Boucard, A.A., Ozkan, E., Strop, P., Newell, E., Sudhof, T.C., Brunger, A.T.(2007) Neuron 56: 992-1003
- PubMed: 18093522 Search on PubMed
- DOI: https://doi.org/10.1016/j.neuron.2007.12.002
- Primary Citation Related Structures:
3BIW, 3BIX - PubMed Abstract:
Neurexins and neuroligins provide trans-synaptic connectivity by the Ca2+-dependent interaction of their alternatively spliced extracellular domains. Neuroligins specify synapses in an activity-dependent manner, presumably by binding to neurexins. Here, we present the crystal structures of neuroligin-1 in isolation and in complex with neurexin-1 beta. Neuroligin-1 forms a constitutive dimer, and two neurexin-1 beta monomers bind to two identical surfaces on the opposite faces of the neuroligin-1 dimer to form a heterotetramer. The neuroligin-1/neurexin-1 beta complex exhibits a nanomolar affinity and includes a large binding interface that contains bound Ca2+. Alternatively spliced sites in neurexin-1 beta and in neuroligin-1 are positioned nearby the binding interface, explaining how they regulate the interaction. Structure-based mutations of neuroligin-1 at the interface disrupt binding to neurexin-1 beta, but not the folding of neuroligin-1 and confirm the validity of the binding interface of the neuroligin-1/neurexin-1 beta complex. Our results provide molecular insights for understanding the role of cell-adhesion proteins in synapse function.
- Howard Hughes Medical Institute, Stanford University, Stanford, CA 94305, USA.
Organizational Affiliation: