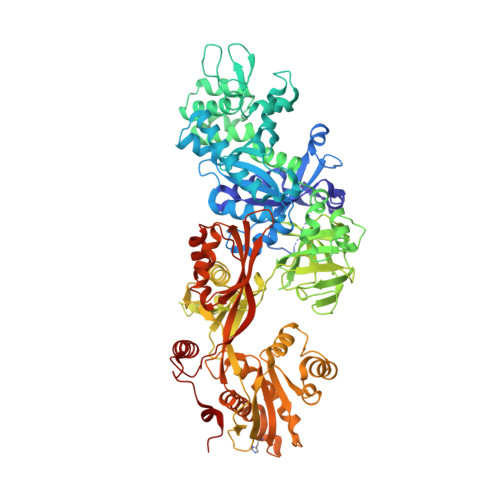
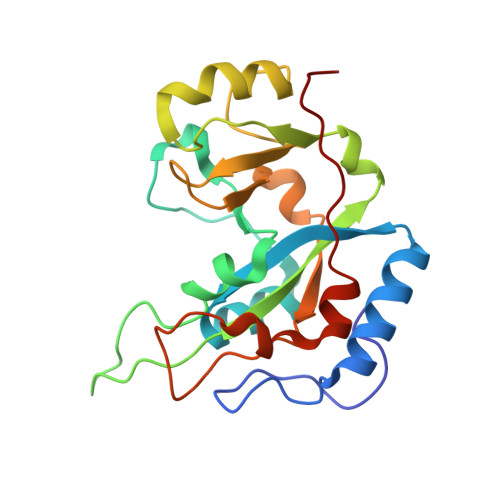
The nature and character of the transition state for the ADP-ribosyltransferase reaction.
Jorgensen, R., Wang, Y., Visschedyk, D., Merrill, A.R.(2008) EMBO Rep 9: 802-809
- PubMed: 18583986 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/embor.2008.90
- Primary Citation Related Structures:
2ZIT, 3B78, 3B82, 3B8H - PubMed Abstract:
Exotoxin A (ExoA) from Pseudomonas aeruginosa is an important virulence factor that belongs to a class of exotoxins that are secreted by pathogenic bacteria which cause human diseases such as cholera, diphtheria, pneumonia and whooping cough. We present the first crystal structures, to our knowledge, of ExoA in complex with elongation factor 2 (eEF2) and intact NAD(+), which indicate a direct role of two active-site loops in ExoA during the catalytic cycle. One loop moves to form a solvent cover for the active site of the enzyme and reaches towards the target residue (diphthamide) in eEF2 forming an important hydrogen bond. The NAD(+) substrate adopts a conformation remarkably different from that of the NAD(+) analogue, betaTAD, observed in previous structures, and fails to trigger any loop movements. Mutational studies of the two loops in the toxin identify several residues important for catalytic activity, in particular Glu 546 and Arg 551, clearly supporting the new complex structures. On the basis of these data, we propose a transition-state model for the toxin-catalysed reaction.
- Department of Molecular and Cellular Biology, University of Guelph, Guelph, Ontario, Canada.
Organizational Affiliation: