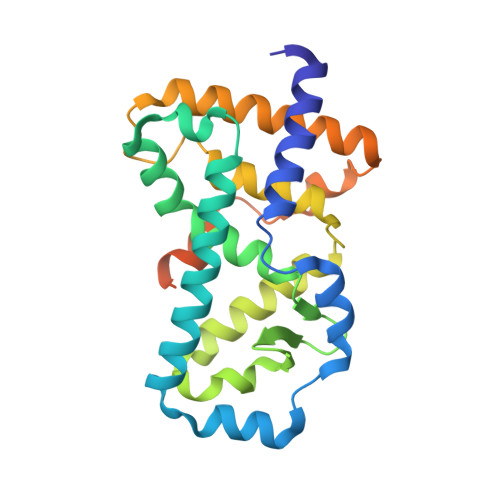
Structural Basis of Digoxin That Antagonizes ROR{gamma}t Receptor Activity and Suppresses Th17 Cell Differentiation and Interleukin (IL)-17 Production
Fujita-Sato, S., Ito, S., Isobe, T., Ohyama, T., Wakabayashi, K., Morishita, K., Ando, O., Isono, F.(2011) J Biological Chem 286: 31409-31417
- PubMed: 21733845 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M111.254003
- Primary Citation Related Structures:
3B0W - PubMed Abstract:
The retinoic acid-related orphan nuclear receptor γt (RORγt)/RORγ2 is well known as a master regulator of interleukin 17 (IL-17)-producing helper T (Th17) cell development. To develop a therapeutic agent against Th17-mediated autoimmune diseases, we screened chemical compounds and successfully found that digoxin inhibited IL-17 production. Further studies revealed that digoxin bound to the ligand binding domain of RORγt and suppressed Th17 differentiation without affecting Th1 differentiation. To better understand the structural basis for the inhibitory activity of digoxin, we determined the crystal structure of the RORγt ligand-binding domain in complex with digoxin at 2.2 Å resolution. The structure reveals that digoxin binds to the ligand-binding pocket protruding between helices H3 and H11 from the pocket. In addition, digoxin disrupts the key interaction important for the agonistic activity, resulting in preventing the positioning of helix H12 in the active conformation, thus antagonizing coactivator interaction. Functional studies demonstrated that digoxin inhibited RORγt activity and decreased IL-17 production but not RORα activity. Digoxin inhibited IL-17 production in CD4(+) T cells from experimental autoimmune encephalomyelitis mice. Our data indicates that RORγt is a promising therapeutic target for Th17-derived autoimmune diseases and our structural data will help to design novel RORγt antagonists.
- Oncology Research Laboratories, R&D Division, Daiichi Sankyo Co., Ltd., Tokyo 134-8630, Japan.
Organizational Affiliation: