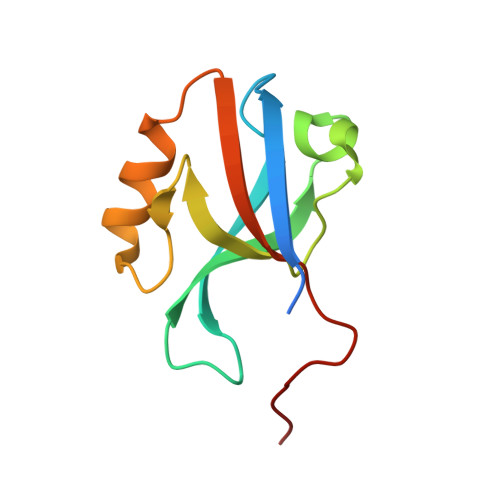
Crystal structure of afadin PDZ domain-nectin-3 complex shows the structural plasticity of the ligand-binding site.
Fujiwara, Y., Goda, N., Tamashiro, T., Narita, H., Satomura, K., Tenno, T., Nakagawa, A., Oda, M., Suzuki, M., Sakisaka, T., Takai, Y., Hiroaki, H.(2015) Protein Sci 24: 376-385
- PubMed: 25534554 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.2628
- Primary Citation Related Structures:
3AXA - PubMed Abstract:
Afadin, a scaffold protein localized in adherens junctions (AJs), links nectins to the actin cytoskeleton. Nectins are the major cell adhesion molecules of AJs. At the initial stage of cell-cell junction formation, the nectin-afadin interaction plays an indispensable role in AJ biogenesis via recruiting and tethering other components. The afadin PDZ domain (AFPDZ) is responsible for binding the cytoplasmic C-terminus of nectins. AFPDZ is a class II PDZ domain member, which prefers ligands containing a class II PDZ-binding motif, X-Φ-X-Φ (Φ, hydrophobic residues); both nectins and other physiological AFPDZ targets contain this class II motif. Here, we report the first crystal structure of the AFPDZ in complex with the nectin-3 C-terminal peptide containing the class II motif. We engineered the nectin-3 C-terminal peptide and AFPDZ to produce an AFPDZ-nectin-3 fusion protein and succeeded in obtaining crystals of this complex as a dimer. This novel dimer interface was created by forming an antiparallel β sheet between β2 strands. A major structural change compared with the known AFPDZ structures was observed in the α2 helix. We found an approximately 2.5 Å-wider ligand-binding groove, which allows the PDZ to accept bulky class II ligands. Apparently, the last three amino acids of the nectin-3 C-terminus were sufficient to bind AFPDZ, in which the two hydrophobic residues are important.
- Division of Structural Biology, Graduate School of Medicine, Kobe University, 7-5-1 Kusunoki-cho, Chuo-ku, Kobe, Hyogo, 650-0017, Japan; Research Center for Structural and Functional Proteomics, Institute for Protein Research, Osaka University, 3-2 Yamadaoka, Suita, 565-0871, Japan; Global-COE (Center of Excellence) Program for Integrative Membrane Biology, Kobe University, 7-5-1 Kusunoki-cho, Chuo, Kobe, Hyogo, 650-0017, Japan.
Organizational Affiliation: