Structural and biochemical characterization of ClfB:ligand interactions
Ganesh, V.K., Barbu, E.M., Deivanayagam, C.C.S., Le, B., Anderson, A., Matsuka, Y., Lin, S.L., Foster, T.J., Narayana, SV.L., Hook, M.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Clumping factor B | 328 | Staphylococcus aureus subsp. aureus N315 | Mutation(s): 1 Gene Names: clfB, SA2423 | 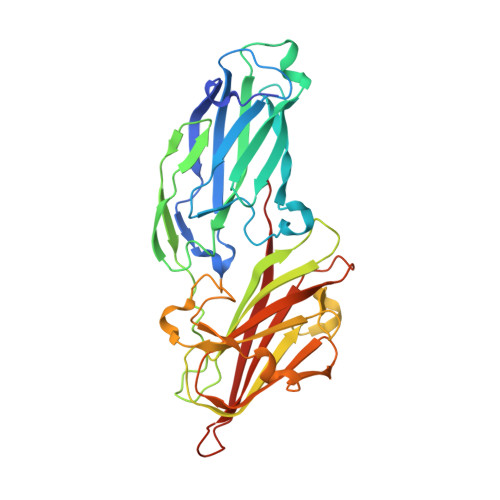 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | Q7A382 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Tail region derived peptide | 15 | Homo sapiens | Mutation(s): 0 | 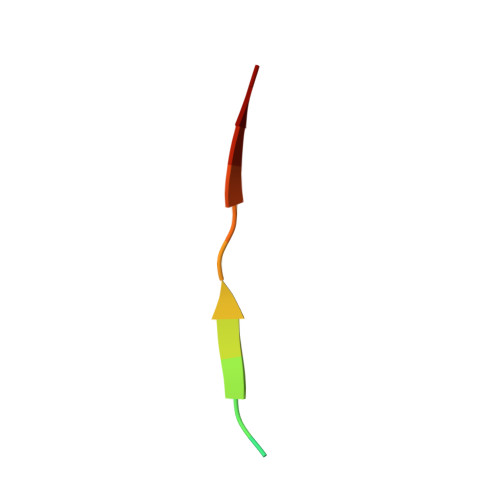 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P13645 GTEx: ENSG00000186395 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P13645 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 85.598 | α = 90 |
| b = 85.598 | β = 90 |
| c = 84.795 | γ = 90 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| CrystalClear | data collection |
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| PHASER | phasing |