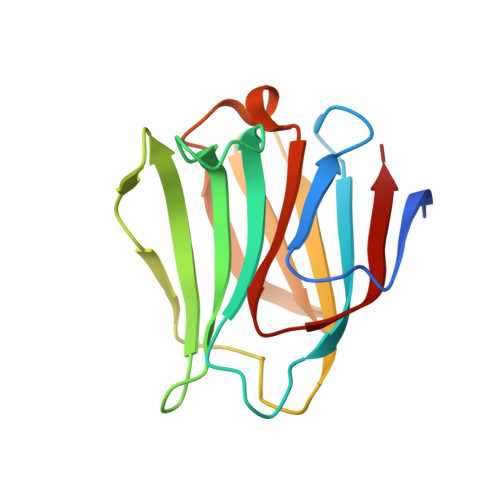
Galectin-8-N-domain recognition mechanism for sialylated and sulfated glycans
Ideo, H., Matsuzaka, T., Nonaka, T., Seko, A., Yamashita, K.(2011) J Biological Chem 286: 11346-11355
- PubMed: 21288902
- DOI: https://doi.org/10.1074/jbc.M110.195925
- Primary Citation of Related Structures:
3AP4, 3AP5, 3AP6, 3AP7, 3AP9 - PubMed Abstract:
Galectin-8 has much higher affinity for 3'-O-sulfated or 3'-O-sialylated glycoconjugates and a Lewis X-containing glycan than for oligosaccharides terminating in Galβ1→3/4GlcNAc, and this specificity is mainly attributed to the N-terminal carbohydrate recognition domain (N-domain, CRD) (Ideo, H., Seko, A., Ishizuka, I., and Yamashita, K. (2003) Glycobiology 13, 713-723). In this study, we elucidated the crystal structures of the human galectin-8-N-domain (-8N) in the absence or presence of 4 ligands. The apo molecule forms a dimer, which is different from the canonical 2-fold symmetric dimer observed for galectin-1 and -2. In a galectin-8N-lactose complex, the lactose-recognizing amino acids are highly conserved among the galectins. However, Arg(45), Gln(47), Arg(59), and the long loop region between the S3 and S4 β-strands are unique to galectin-8N. These amino acids directly or indirectly interact with the sulfate or sialic acid moieties of 3'-sialyl- and 3'-sulfolactose complexed with galectin-8N. Furthermore, in the LNF-III-galectin-8N complex, van der Waals interactions occur between the α1-3-branched fucose and galactose and between galactose and Tyr(141), and these interactions increase the affinity toward galectin-8N. Based on the findings of these x-ray crystallographic analyses, a mutagenesis study using surface plasmon resonance showed that Arg(45), Gln(47), and Arg(59) of galectin-8N are indispensable and coordinately contribute to the strong binding of galectins-8N to sialylated and sulfated oligosaccharides. Arg(59) is the most critical amino acid for binding in the S3-S4 loop region.
- Innovative Research Initiatives, Tokyo Institute of Technology, Yokohama, Japan.
Organizational Affiliation: