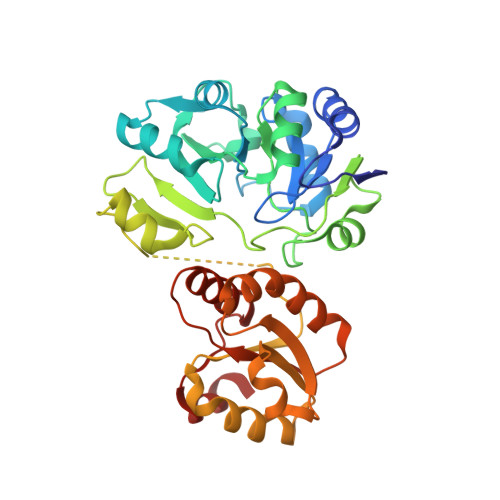
Crystal structure of Sulfolobus tokodaii Sua5 complexed with L-threonine and AMPPNP
Kuratani, M., Kasai, T., Akasaka, R., Higashijima, K., Terada, T., Kigawa, T., Shinkai, A., Bessho, Y., Yokoyama, S.(2011) Proteins 79: 2065-2075
- PubMed: 21538543 Search on PubMed
- DOI: https://doi.org/10.1002/prot.23026
- Primary Citation Related Structures:
3AJE - PubMed Abstract:
The hypermodified nucleoside N(6)-threonylcarbamoyladenosine resides at position 37 of tRNA molecules bearing U at position 36 and maintains translational fidelity in the three kingdoms of life. The N(6)-threonylcarbamoyl moiety is composed of L-threonine and bicarbonate, and its synthesis was genetically shown to require YrdC/Sua5. YrdC/Sua5 binds to tRNA and ATP. In this study, we analyzed the L-threonine-binding mode of Sua5 from the archaeon Sulfolobus tokodaii. Isothermal titration calorimetry measurements revealed that S. tokodaii Sua5 binds L-threonine more strongly than L-serine and glycine. The Kd values of Sua5 for L-threonine and L-serine are 9.3 μM and 2.6 mM, respectively. We determined the crystal structure of S. tokodaii Sua5, complexed with AMPPNP and L-threonine, at 1.8 Å resolution. The L-threonine is bound next to AMPPNP in the same pocket of the N-terminal domain. Thr118 and two water molecules form hydrogen bonds with AMPPNP in a unique manner for adenine-specific recognition. The carboxyl group and the side-chain hydroxyl and methyl groups of L-threonine are buried deep in the pocket, whereas the amino group faces AMPPNP. The L-threonine is located in a suitable position to react together with ATP for the synthesis of N(6)-threonylcarbamoyladenosine.
- RIKEN Systems and Structural Biology Center, Yokohama 230-0045, Japan.
Organizational Affiliation: