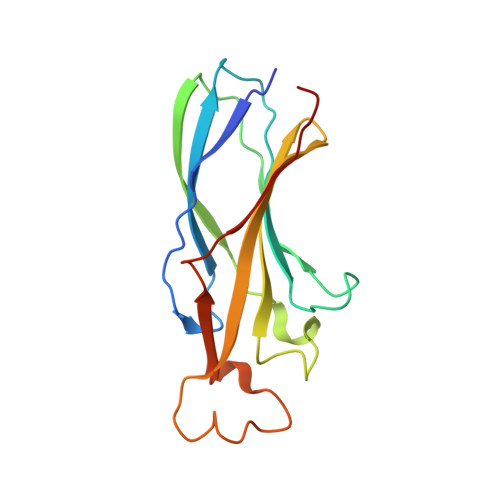
Structure of the histone chaperone CIA/ASF1-double bromodomain complex linking histone modifications and site-specific histone eviction
Akai, Y., Adachi, N., Hayashi, Y., Eitoku, M., Sano, N., Natsume, R., Kudo, N., Tanokura, M., Senda, T., Horikoshi, M.(2010) Proc Natl Acad Sci U S A 107: 8153-8158
- PubMed: 20393127 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0912509107
- Primary Citation Related Structures:
3AAD - PubMed Abstract:
Nucleosomes around the promoter region are disassembled for transcription in response to various signals, such as acetylation and methylation of histones. Although the interactions between histone-acetylation-recognizing bromodomains and factors involved in nucleosome disassembly have been reported, no structural basis connecting histone modifications and nucleosome disassembly has been obtained. Here, we determined at 3.3 A resolution the crystal structure of histone chaperone cell cycle gene 1 (CCG1) interacting factor A/antisilencing function 1 (CIA/ASF1) in complex with the double bromodomain in the CCG1/TAF1/TAF(II)250 subunit of transcription factor IID. Structural, biochemical, and biological studies suggested that interaction between double bromodomain and CIA/ASF1 is required for their colocalization, histone eviction, and pol II entry at active promoter regions. Furthermore, the present crystal structure has characteristics that can connect histone acetylation and CIA/ASF1-mediated histone eviction. These findings suggest that the molecular complex between CIA/ASF1 and the double bromodomain plays a key role in site-specific histone eviction at active promoter regions. The model we propose here is the initial structure-based model of the biological signaling from histone modifications to structural change of the nucleosome (hi-MOST model).
- Biomedicinal Information Research Center, National Institute of Advanced Industrial Science and Technology, 2-4-7 Aomi, Koto-ku, Tokyo 135-0064, Japan.
Organizational Affiliation: