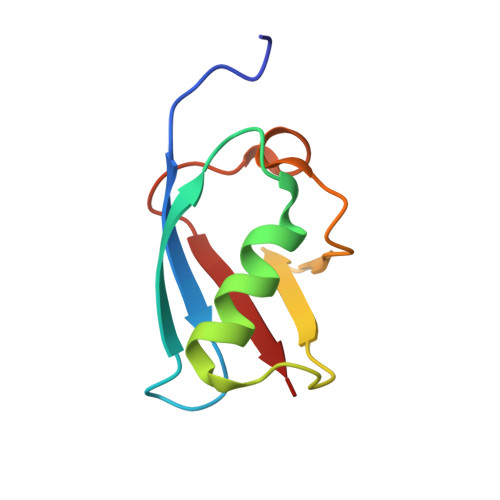
Structural basis for regulation of poly-SUMO chain by a SUMO-like domain of Nip45
Sekiyama, N., Arita, K., Ikeda, Y., Hashiguchi, K., Ariyoshi, M., Tochio, H., Saitoh, H., Shirakawa, M.(2009) Proteins 78: 1491-1502
- PubMed: 20077568 Search on PubMed
- DOI: https://doi.org/10.1002/prot.22667
- Primary Citation Related Structures:
3A4R, 3A4S - PubMed Abstract:
Post-translational modification by small ubiquitin-like modifier (SUMO) provides an important regulatory mechanism in diverse cellular processes. Modification of SUMO has been shown to target proteins involved in systems ranging from DNA repair pathways to the ubiquitin-proteasome degradation system by the action of SUMO-targeted ubiquitin ligases (STUbLs). STUbLs recognize target proteins modified with a poly-SUMO chain through their SUMO-interacting motifs (SIMs). STUbLs are also associated with RENi family proteins, which commonly have two SUMO-like domains (SLD1 and SLD2) at their C terminus. We have determined the crystal structures of SLD2 of mouse RENi protein, Nip45, in a free form and in complex with a mouse E2 sumoylation enzyme, Ubc9. While Nip45 SLD2 shares a beta-grasp fold with SUMO, the SIM interaction surface conserved in SUMO paralogues does not exist in SLD2. Biochemical data indicates that neither tandem SLDs or SLD2 of Nip45 bind to either tandem SIMs from either mouse STUbL, RNF4 or to those from SUMO-binding proteins, whose interactions with SUMO have been well characterized. On the other hand, Nip45 SLD2 binds to Ubc9 in an almost identical manner to that of SUMO and thereby inhibits elongation of poly-SUMO chains. This finding highlights a possible role of the RENi proteins in the modulation of Ubc9-mediated poly-SUMO formation.
- Kyoto University, Japan.
Organizational Affiliation: