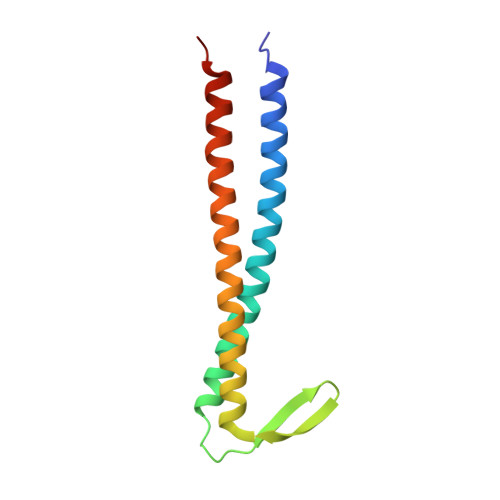
Structural and molecular characterization of the prefoldin beta subunit from Thermococcus strain KS-1
Kida, H., Sugano, Y., Iizuka, R., Fujihashi, M., Yohda, M., Miki, K.(2008) J Mol Biology 383: 465-474
- PubMed: 18775436
- DOI: https://doi.org/10.1016/j.jmb.2008.08.041
- Primary Citation of Related Structures:
2ZQM - PubMed Abstract:
Prefoldin (PFD) is a heterohexameric molecular chaperone that is found in eukaryotic cytosol and archaea. PFD is composed of alpha and beta subunits and forms a "jellyfish-like" structure. PFD binds and stabilizes nascent polypeptide chains and transfers them to group II chaperonins for completion of their folding. Recently, the whole genome of Thermococcus kodakaraensis KOD1 was reported and shown to contain the genes of two alpha and two beta subunits of PFD. The genome of Thermococcus strain KS-1 also possesses two sets of alpha (alpha1 and alpha2) and beta subunits (beta1 and beta2) of PFD (TsPFD). However, the functions and roles of each of these PFD subunits have not been investigated in detail. Here, we report the crystal structure of the TsPFD beta1 subunit at 1.9 A resolution and its functional analysis. TsPFD beta1 subunits form a tetramer with four coiled-coil tentacles resembling the jellyfish-like structure of heterohexameric PFD. The beta hairpin linkers of beta1 subunits assemble to form a beta barrel "body" around a central fourfold axis. Size-exclusion chromatography and multi-angle light-scattering analyses show that the beta1 subunits form a tetramer at pH 8.0 and a dimer of tetramers at pH 6.8. The tetrameric beta1 subunits can protect against aggregation of relatively small proteins, insulin or lysozyme. The structural and biochemical analyses imply that PFD beta1 subunits act as molecular chaperones in living cells of some archaea.
- Department of Chemistry, Graduate School of Science, Kyoto University, Sakyo-ku, Kyoto, 606-8502, Japan.
Organizational Affiliation: