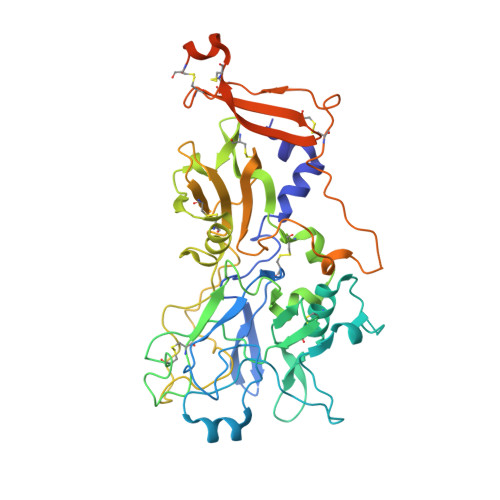
Structural Characterization of Apical Membrane Antigen 1 (Ama1) from Toxoplasma Gondii.
Crawford, J., Tonkin, M.L., Grujic, O., Boulanger, M.J.(2010) J Biological Chem 285: 15644
- PubMed: 20304917 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M109.092619
- Primary Citation Related Structures:
2X2Z - PubMed Abstract:
Apical membrane antigen 1 (AMA1) is an essential component of the moving junction complex used by Apicomplexan parasites to invade host cells. We report the 2.0 A resolution x-ray crystal structure of the full ectodomain (domains I, II, and III) of AMA1 from the pervasive protozoan parasite Toxoplasma gondii. The structure of T. gondii AMA1 (TgAMA1) is the most complete of any AMA1 structure to date, with more than 97.5% of the ectodomain unambiguously modeled. Comparative sequence analysis reveals discrete segments of divergence in TgAMA1 that map to areas of established functional importance in AMA1 from Plasmodium vivax (PvAMA1) and Plasmodium falciparum (PfAMA1). Inspection of the TgAMA1 structure reveals a network of apical surface loops, reorganized in both size and chemistry relative to PvAMA1/PfAMA1, that appear to serve as structural filters restricting access to a central hydrophobic groove. The terminal portion of this groove is formed by an extended loop from DII that is 14 residues shorter in TgAMA1. A pair of tryptophan residues (Trp(353) and Trp(354)) anchor the DII loop in the hydrophobic groove and frame a conserved tyrosine (Tyr(230)), forming a contiguous surface that may be critical for moving junction assembly. The minimalist DIII structure folds into a cystine knot that probably stabilizes and orients the bulk of the ectodmain without providing excess surface area to which invasion-inhibitory antibodies can be generated. The detailed structural characterization of TgAMA1 provides valuable insight into the mechanism of host cell invasion by T. gondii.
- Department of Biochemistry and Microbiology, University of Victoria, Victoria, British Columbia V8W 3P6, Canada.
Organizational Affiliation: