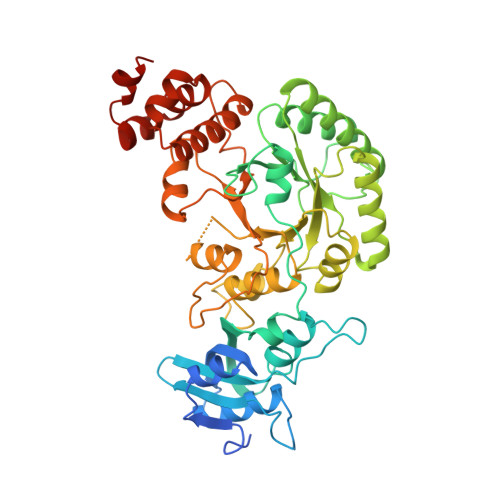
Crystal structure of the ternary complex of ribulose-1,5-bisphosphate carboxylase, Mg(II), and activator CO2 at 2.3-A resolution.
Lundqvist, T., Schneider, G.(1991) Biochemistry 30: 904-908
- PubMed: 1899197 Search on PubMed
- DOI: https://doi.org/10.1021/bi00218a004
- Primary Citation Related Structures:
2RUS - PubMed Abstract:
The activated ternary complex, enzyme-CO2-Mg(II), of the dimeric ribulose-1,5-bisphosphate carboxylase/oxygenase from Rhodospirillum rubrum can be prepared in the same crystal form that was used for the crystallographic structure determination of the native nonactivated enzyme (Schneider, G., Bränden, C.-I., & Lorimer, G. (1986) J. Mol. Biol. 187, 141-143). The three-dimensional structure of the activated enzyme has been determined to a nominal resolution of 2.3 A by protein crystallographic methods. The activator CO2 forms a carbamate with Lys191, located at the bottom of the funnel-shaped active site. In both subunits, this labile adduct is stabilized by a Mg(II) ion, bound to the carbamate and the side chains of Asp193 and Glu194. One solvent molecule was found within the first coordination sphere of the metal ion. The metal-binding site in ribulose-1,5-bisphosphate carboxylase consists thus of at least three protein ligands, all located on loop 2 of the beta/alpha barrel. One additional metal ligand, the side chain of the conserved Asn111, was observed close to the Mg(II) ion in the B-subunit. Other structural differences at the active site between the activated and nonactivated enzyme are limited to side-chain positions. Nevertheless, it is obvious that the hydrogen-bonding pattern in the vicinity of the activator site is completely altered.
- Department of Molecular Biology, Swedish University of Agricultural Sciences, Uppsala.
Organizational Affiliation: