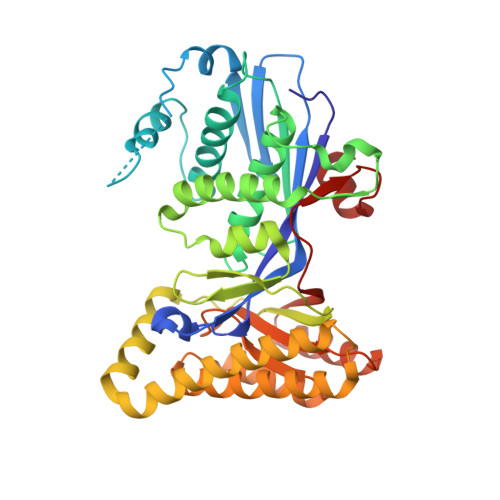
Biochemical and Structural Basis for Feedback Inhibition of Mevalonate Kinase and Isoprenoid Metabolism.
Fu, Z., Voynova, N.E., Herdendorf, T.J., Miziorko, H.M., Kim, J.J.(2008) Biochemistry 47: 3715-3724
- PubMed: 18302342 Search on PubMed
- DOI: https://doi.org/10.1021/bi7024386
- Primary Citation Related Structures:
2R3V, 2R42 - PubMed Abstract:
Mevalonate kinase (MK), which catalyzes a key reaction in polyisoprenoid and sterol metabolism in many organisms, is subject to feedback regulation by farnesyl diphosphate and related compounds. The structures of human mevalonate kinase and a binary complex of the rat enzyme incubated with farnesyl thiodiphosphate (FSPP) are reported. Significant FSPP hydrolysis occurs under crystallization conditions; this results in detection of farnesyl thiophosphate (FSP) in the structure of the binary complex. Farnesyl thiodiphosphate competes with substrate ATP to produce feedback inhibition of mevalonate kinase. The binding sites for these metabolites overlap, with the phosphate of FSP nearly superimposed on ATP's beta-phosphate and FSP's polyisoprenoid chain overlapping ATP's adenosine moiety. Several hydrophobic amino acid side chains are positioned near the polyisoprenoid chain of FSP and their functional significance has been evaluated in mutagenesis experiments with human MK, which exhibits the highest reported sensitivity to feedback inhibition. Results suggest that single and double mutations at T104 and I196 produce a significant inflation of the K(i) for FSPP (approximately 40-fold for T104A/I196A). Such an effect persists when K(i) values are normalized for effects on the K(m) for ATP, suggesting that it may be possible to engineer MK proteins with altered sensitivity to feedback inhibition. Comparison of animal MK protein alignments and structures with those of a MK protein from Streptococcus pneumoniae indicates that sequence differences between N- and C-terminal domains correlate with differences in interdomain angles. Bacterial MK proteins exhibit more solvent exposure of feedback inhibitor binding sites and, consequently, weaker binding of these inhibitors.
- Department of Biochemistry, Medical College of Wisconsin, Milwaukee, Wisconsin 53226, USA.
Organizational Affiliation: