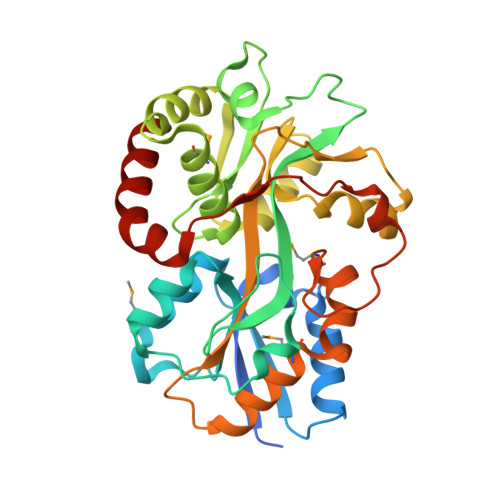
Structural Similarities between Thiamin-Binding Protein and Thiaminase-I Suggest a Common Ancestor
Soriano, E.V., Rajashankar, K.R., Hanes, J.W., Bale, S., Begley, T.P., Ealick, S.E.(2008) Biochemistry 47: 1346-1357
- PubMed: 18177053
- DOI: https://doi.org/10.1021/bi7018282
- Primary Citation of Related Structures:
2QRY - PubMed Abstract:
ATP-binding cassette (ABC) transporters are responsible for the transport of a wide variety of water-soluble molecules and ions into prokaryotic cells. In Gram-negative bacteria, periplasmic-binding proteins deliver ions or molecules such as thiamin to the membrane-bound ABC transporter. The gene for the thiamin-binding protein tbpA has been identified in both Escherichia coli and Salmonella typhimurium. Here we report the crystal structure of TbpA from E. coli with bound thiamin monophosphate. The structure was determined at 2.25 A resolution using single-wavelength anomalous diffraction experiments, despite the presence of nonmerohedral twinning. The crystal structure shows that TbpA belongs to the group II periplasmic-binding protein family. Equilibrium binding measurements showed similar dissociation constants for thiamin, thiamin monophosphate, and thiamin pyrophosphate. Analysis of the binding site by molecular modeling demonstrated how TbpA binds all three forms of thiamin. A comparison of TbpA and thiaminase-I, a thiamin-degrading enzyme, revealed structural similarity between the two proteins, especially in domain 1, suggesting that the two proteins evolved from a common ancestor.
- Department of Chemistry and Chemical Biology, Cornell University, Ithaca, New York 14853, USA.
Organizational Affiliation: