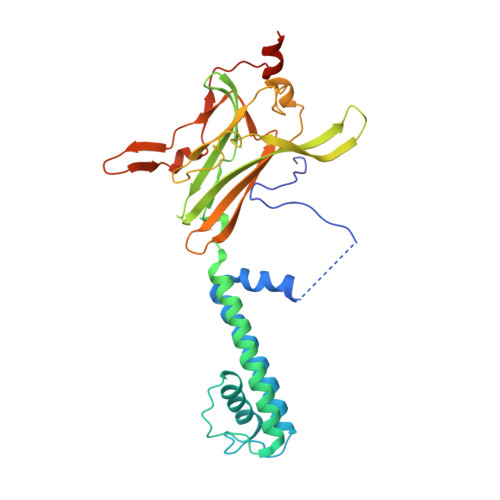
Crystal structure of a Kir3.1-prokaryotic Kir channel chimera.
Nishida, M., Cadene, M., Chait, B.T., Mackinnon, R.(2007) EMBO J 26: 4005-4015
- PubMed: 17703190 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7601828
- Primary Citation Related Structures:
2QKS - PubMed Abstract:
The Kir3.1 K(+) channel participates in heart rate control and neuronal excitability through G-protein and lipid signaling pathways. Expression in Escherichia coli has been achieved by replacing three fourths of the transmembrane pore with the pore of a prokaryotic Kir channel, leaving the cytoplasmic pore and membrane interfacial regions of Kir3.1 origin. Two structures were determined at 2.2 A. The selectivity filter is identical to the Streptomyces lividans K(+) channel within error of measurement (r.m.s.d.<0.2 A), suggesting that K(+) selectivity requires extreme conservation of three-dimensional structure. Multiple K(+) ions reside within the pore and help to explain voltage-dependent Mg(2+) and polyamine blockade and strong rectification. Two constrictions, at the inner helix bundle and at the apex of the cytoplasmic pore, may function as gates: in one structure the apex is open and in the other, it is closed. Gating of the apex is mediated by rigid-body movements of the cytoplasmic pore subunits. Phosphatidylinositol 4,5-biphosphate-interacting residues suggest a possible mechanism by which the signaling lipid regulates the cytoplasmic pore.
- Laboratory of Molecular Neurobiology and Biophysics, Howard Hughes Medical Institute, Rockefeller University, New York, NY 10065, USA.
Organizational Affiliation: