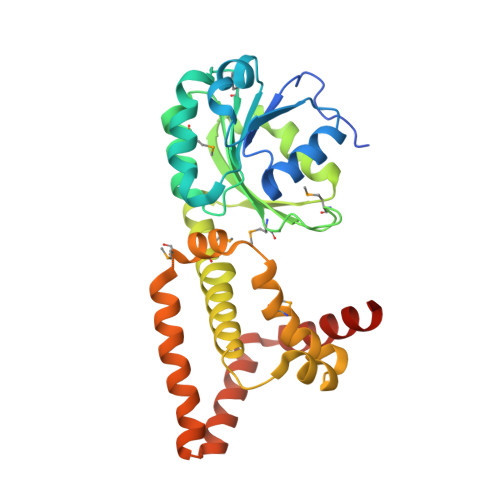
The structure of Haemophilus influenzae prephenate dehydrogenase suggests unique features of bifunctional TyrA enzymes.
Chiu, H.J., Abdubek, P., Astakhova, T., Axelrod, H.L., Carlton, D., Clayton, T., Das, D., Deller, M.C., Duan, L., Feuerhelm, J., Grant, J.C., Grzechnik, A., Han, G.W., Jaroszewski, L., Jin, K.K., Klock, H.E., Knuth, M.W., Kozbial, P., Krishna, S.S., Kumar, A., Marciano, D., McMullan, D., Miller, M.D., Morse, A.T., Nigoghossian, E., Okach, L., Reyes, R., Tien, H.J., Trame, C.B., van den Bedem, H., Weekes, D., Xu, Q., Hodgson, K.O., Wooley, J., Elsliger, M.A., Deacon, A.M., Godzik, A., Lesley, S.A., Wilson, I.A.(2010) Acta Crystallogr Sect F Struct Biol Cryst Commun 66: 1317-1325
- PubMed: 20944228 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S1744309110021688
- Primary Citation Related Structures:
2PV7 - PubMed Abstract:
Chorismate mutase/prephenate dehydrogenase from Haemophilus influenzae Rd KW20 is a bifunctional enzyme that catalyzes the rearrangement of chorismate to prephenate and the NAD(P)(+)-dependent oxidative decarboxylation of prephenate to 4-hydroxyphenylpyruvate in tyrosine biosynthesis. The crystal structure of the prephenate dehydrogenase component (HinfPDH) of the TyrA protein from H. influenzae Rd KW20 in complex with the inhibitor tyrosine and cofactor NAD(+) has been determined to 2.0 Å resolution. HinfPDH is a dimeric enzyme, with each monomer consisting of an N-terminal α/β dinucleotide-binding domain and a C-terminal α-helical dimerization domain. The structure reveals key active-site residues at the domain interface, including His200, Arg297 and Ser179 that are involved in catalysis and/or ligand binding and are highly conserved in TyrA proteins from all three kingdoms of life. Tyrosine is bound directly at the catalytic site, suggesting that it is a competitive inhibitor of HinfPDH. Comparisons with its structural homologues reveal important differences around the active site, including the absence of an α-β motif in HinfPDH that is present in other TyrA proteins, such as Synechocystis sp. arogenate dehydrogenase. Residues from this motif are involved in discrimination between NADP(+) and NAD(+). The loop between β5 and β6 in the N-terminal domain is much shorter in HinfPDH and an extra helix is present at the C-terminus. Furthermore, HinfPDH adopts a more closed conformation compared with TyrA proteins that do not have tyrosine bound. This conformational change brings the substrate, cofactor and active-site residues into close proximity for catalysis. An ionic network consisting of Arg297 (a key residue for tyrosine binding), a water molecule, Asp206 (from the loop between β5 and β6) and Arg365' (from the additional C-terminal helix of the adjacent monomer) is observed that might be involved in gating the active site.
- Stanford Synchrotron Radiation Lightsource, SLAC National Accelerator Laboratory, Menlo Park, CA, USA.
Organizational Affiliation: