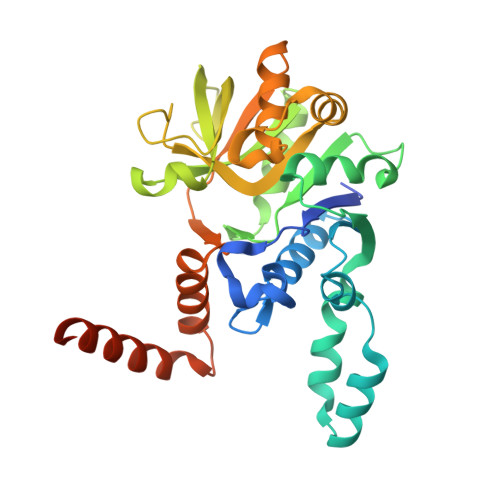
Active site geometry of glucose-1-phosphate uridylyltransferase.
Thoden, J.B., Holden, H.M.(2007) Protein Sci 16: 1379-1388
- PubMed: 17567737 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.072864707
- Primary Citation Related Structures:
2PA4 - PubMed Abstract:
Glucose-1-phosphate uridylyltransferase, or UGPase, catalyzes the production of UDP-glucose from glucose-1-phosphate and UTP. Because of the biological role of UDP-glucose in glycogen synthesis and in the formation of glycolipids, glycoproteins, and proteoglycans, the enzyme is widespread in nature. Recently this laboratory reported the three-dimensional structure of UGPase from Escherichia coli. While the initial X-ray analysis revealed the overall fold of the enzyme, details concerning its active site geometry were limited because crystals of the protein complexed with either substrates or products could never be obtained. In an effort to more fully investigate the active site geometry of the enzyme, UGPase from Corynebacterium glutamicum was subsequently cloned and purified. Here we report the X-ray structure of UGPase crystallized in the presence of both magnesium and UDP-glucose. Residues involved in anchoring the ligand to the active site include the polypeptide chain backbone atoms of Ala 20, Gly 21, Gly 117, Gly 180, and Ala 214, and the side chains of Glu 36, Gln 112, Asp 143, Glu 201, and Lys 202. Two magnesium ions are observed coordinated to the UDP-glucose. An alpha- and a beta-phosphoryl oxygen, three waters, and the side chain of Asp 142 ligate the first magnesium, whereas the second ion is coordinated by an alpha-phosphoryl oxygen and five waters. The position of the first magnesium is conserved in both the glucose-1-phosphate thymidylyltransferases and the cytidylyltransferases. The structure presented here provides further support for the role of the conserved magnesium ion in the catalytic mechanisms of the sugar-1-phosphate nucleotidylyltransferases.
- Department of Biochemistry, University of Wisconsin, Madison 53706-1544, USA.
Organizational Affiliation: